INDETERMINATE1-mediated expression of FT family genes is required for proper timing of flowering in Brachypodium distachyon
- PMID: 37934817
- PMCID: PMC10655584
- DOI: 10.1073/pnas.2312052120
INDETERMINATE1-mediated expression of FT family genes is required for proper timing of flowering in Brachypodium distachyon
Abstract
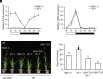
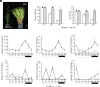
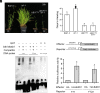
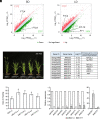
The transition to flowering is a major developmental switch in plants. In many temperate grasses, perception of indicators of seasonal change, such as changing day-length and temperature, leads to expression of FLOWERING LOCUS T1 (FT1) and FT-Like (FTL) genes that are essential for promoting the transition to flowering. However, little is known about the upstream regulators of FT1 and FTL genes in temperate grasses. Here, we characterize the monocot-specific gene INDETERMINATE1 (BdID1) in Brachypodium distachyon and demonstrate that BdID1 is a regulator of FT family genes. Mutations in ID1 impact the ability of the short-day (SD) vernalization, cold vernalization, and long-day (LD) photoperiod pathways to induce certain FTL genes. BdID1 is required for upregulation of FTL9 (FT-LIKE9) expression by the SD vernalization pathway, and overexpression of FTL9 in an id1 background can partially restore the delayed flowering phenotype of id1. We show that BdID1 binds in vitro to the promoter region of FTL genes suggesting that ID1 directly activates FTL expression. Transcriptome analysis shows that BdID1 is required for FT1, FT2, FTL12, and FTL13 expression under inductive LD photoperiods, indicating that BdID1 is a regulator of the FT gene family. Moreover, overexpression of FT1 in the id1 background results in rapid flowering similar to overexpressing FT1 in the wild type, demonstrating that BdID1 is upstream of FT family genes. Interestingly, ID1 negatively regulates a previously uncharacterized FTL gene, FTL4, and we show that FTL4 is a repressor of flowering. Thus, BdID1 is critical for proper timing of flowering in temperate grasses.
Keywords: Brachypodium distachyon; FT; INDETERMINATE1; flowering; vernalization.
Conflict of interest statement
The authors declare no competing interest.
Figures


Similar articles
-
Divergent roles of FT-like 9 in flowering transition under different day lengths in Brachypodium distachyon.Nat Commun. 2019 Feb 18;10(1):812. doi: 10.1038/s41467-019-08785-y. Nat Commun. 2019. PMID: 30778068 Free PMC article.
-
A florigen paralog is required for short-day vernalization in a pooid grass.Elife. 2019 Jan 8;8:e42153. doi: 10.7554/eLife.42153. Elife. 2019. PMID: 30618375 Free PMC article.
-
Characterization of FLOWERING LOCUS T1 (FT1) gene in Brachypodium and wheat.PLoS One. 2014 Apr 9;9(4):e94171. doi: 10.1371/journal.pone.0094171. eCollection 2014. PLoS One. 2014. PMID: 24718312 Free PMC article.
-
The molecular biology of seasonal flowering-responses in Arabidopsis and the cereals.Ann Bot. 2009 Jun;103(8):1165-72. doi: 10.1093/aob/mcp063. Epub 2009 Mar 21. Ann Bot. 2009. PMID: 19304997 Free PMC article. Review.
-
The molecular basis of vernalization in different plant groups.Cold Spring Harb Symp Quant Biol. 2012;77:105-15. doi: 10.1101/sqb.2013.77.014449. Epub 2013 Apr 25. Cold Spring Harb Symp Quant Biol. 2012. PMID: 23619014 Review.
Cited by
-
Functional divergence of FTL9 and FTL10 in flowering control in rice.BMC Genomics. 2024 Jun 5;25(1):562. doi: 10.1186/s12864-024-10441-9. BMC Genomics. 2024. PMID: 38840036 Free PMC article.
-
INDETERMINATE DOMAIN Transcription Factors in Crops: Plant Architecture, Disease Resistance, Stress Response, Flowering, and More.Int J Mol Sci. 2024 Sep 24;25(19):10277. doi: 10.3390/ijms251910277. Int J Mol Sci. 2024. PMID: 39408609 Free PMC article. Review.
-
Unraveling the Complexities of Flowering in Ornamental Plants: The Interplay of Genetics, Hormonal Networks, and Microbiome.Plants (Basel). 2025 Apr 6;14(7):1131. doi: 10.3390/plants14071131. Plants (Basel). 2025. PMID: 40219203 Free PMC article. Review.
-
The H3K4 demethylase JMJ1 is required for proper timing of flowering in Brachypodium distachyon.Plant Cell. 2024 Jul 2;36(7):2729-2745. doi: 10.1093/plcell/koae124. Plant Cell. 2024. PMID: 38652680 Free PMC article.
References
-
- Andres F., Coupland G., The genetic basis of flowering responses to seasonal cues. Nat. Rev. Genet. 13, 627–639 (2012). - PubMed
-
- Chouard P., Vernalization and its relations to dormancy. Annu. Rev. Plant Physiol. Plant Mol. Biol. 11, 191–238 (1960).
-
- Purvis O. N., Gregory F. G., Studies in vernalisation of cereals I. A comparative study of vernalisation of winter rye by low temperature and by short days. Ann. Bot. Lond. 1, 562–569 (1937).
-
- Heide O. M., Control of flowering and reproduction in temperate grasses. New Phytol. 128, 347–362 (1994). - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

