Exploring the role of SWI/SNF complex subunit BAF60c in lipid metabolism and inflammation in fish
- PMID: 37942006
- PMCID: PMC10628743
- DOI: 10.1016/j.isci.2023.108207
Exploring the role of SWI/SNF complex subunit BAF60c in lipid metabolism and inflammation in fish
Abstract
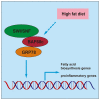
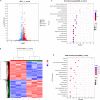
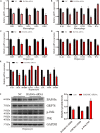
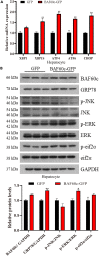
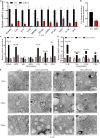
Chromatin remodeling plays an important role in regulating gene transcription, in which chromatin remodeling complex is a crucial aspect. Brg1/Brm-associated factor 60c (BAF60c) subunit forms a bridge between chromatin remodeling complexes and transcription factors in mammals; hence, it has received extensive attention. However, the roles of BAF60c in fish remain largely unexplored. In this study, we identified BAF60c-interacting proteins by using HIS-pull-down and LC-MS/MS analysis in fish. Subsequently, the RNA-seq analysis was performed to identify the overall effects of BAF60c. Then, the function of BAF60c was verified through BAF60c knockdown and overexpression experiments. We demonstrated for the first time that BAF60c interacts with glucose-regulated protein 78 (GRP78) and regulates lipid metabolism, endoplasmic reticulum (ER) stress, and inflammation. Knockdown of BAF60c reduces fatty acid biosynthesis, ER stress, and inflammation. In conclusion, the results enriched BAF60c-interacting protein network and explored the function of BAF60c in lipid metabolism and inflammation in fish.
Keywords: Biological sciences; Molecular biology; Physiology; Zoology.
© 2023 The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures
Similar articles
-
Baf60c in skeletal muscle regulates adipose tissue thermogenesis via Musclin-mediated endocrine signaling.Life Metab. 2025 May 4;4(4):loaf015. doi: 10.1093/lifemeta/loaf015. eCollection 2025 Aug. Life Metab. 2025. PMID: 40585527 Free PMC article.
-
Coordinate Nodal and BMP inhibition directs Baf60c-dependent cardiomyocyte commitment.Genes Dev. 2013 Nov 1;27(21):2332-44. doi: 10.1101/gad.225144.113. Genes Dev. 2013. PMID: 24186978 Free PMC article.
-
The BAF60c-MyoD complex poises chromatin for rapid transcription.Bioarchitecture. 2012 May 1;2(3):104-109. doi: 10.4161/bioa.20970. Bioarchitecture. 2012. PMID: 22880151 Free PMC article.
-
Role of BAF60a/BAF60c in chromatin remodeling and hepatic lipid metabolism.Nutr Metab (Lond). 2016 Apr 27;13:30. doi: 10.1186/s12986-016-0090-1. eCollection 2016. Nutr Metab (Lond). 2016. PMID: 27127533 Free PMC article. Review.
-
The SWI/SNF chromatin-remodeling factors BAF60a, b, and c in nutrient signaling and metabolic control.Protein Cell. 2018 Feb;9(2):207-215. doi: 10.1007/s13238-017-0442-2. Epub 2017 Jul 7. Protein Cell. 2018. PMID: 28688083 Free PMC article. Review.
Cited by
-
The SWI/SNF chromatin remodeling complex: a critical regulator of metabolism.Biochem Soc Trans. 2024 Jun 26;52(3):1327-1337. doi: 10.1042/BST20231141. Biochem Soc Trans. 2024. PMID: 38666605 Free PMC article. Review.
References
LinkOut - more resources
Full Text Sources
Miscellaneous

