A drug-like molecule engages nuclear hormone receptor DAF-12/FXR to regulate mitophagy and extend lifespan
- PMID: 37957360
- PMCID: PMC10797806
- DOI: 10.1038/s43587-023-00524-9
A drug-like molecule engages nuclear hormone receptor DAF-12/FXR to regulate mitophagy and extend lifespan
Abstract
Autophagy-lysosomal function is crucial for maintaining healthy lifespan and preventing age-related diseases. The transcription factor TFEB plays a key role in regulating this pathway. Decreased TFEB expression is associated with various age-related disorders, making it a promising therapeutic target. In this study, we screened a natural product library and discovered mitophagy-inducing coumarin (MIC), a benzocoumarin compound that enhances TFEB expression and lysosomal function. MIC robustly increases the lifespan of Caenorhabditis elegans in an HLH-30/TFEB-dependent and mitophagy-dependent manner involving DCT-1/BNIP3 while also preventing mitochondrial dysfunction in mammalian cells. Mechanistically, MIC acts by inhibiting ligand-induced activation of the nuclear hormone receptor DAF-12/FXR, which, in turn, induces mitophagy and extends lifespan. In conclusion, our study uncovers MIC as a promising drug-like molecule that enhances mitochondrial function and extends lifespan by targeting DAF-12/FXR. Furthermore, we discovered DAF-12/FXR as a previously unknown upstream regulator of HLH-30/TFEB and mitophagy.
© 2023. The Author(s), under exclusive licence to Springer Nature America, Inc.
Figures


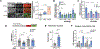



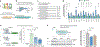



References
-
- Abdrakhmanov A, Gogvadze V. & Zhivotovsky B. To eat or to die: deciphering selective forms of autophagy. Trends Biochem. Sci 45, 347–364 (2020). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

