An Integrated Analysis of microRNAs and the Transcriptome Reveals the Molecular Mechanisms Underlying the Regulation of Leaf Development in Xinyang Maojian Green Tea (Camellia sinensis)
- PMID: 37960023
- PMCID: PMC10649745
- DOI: 10.3390/plants12213665
An Integrated Analysis of microRNAs and the Transcriptome Reveals the Molecular Mechanisms Underlying the Regulation of Leaf Development in Xinyang Maojian Green Tea (Camellia sinensis)
Abstract
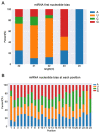
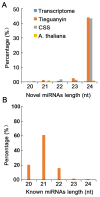
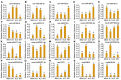
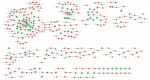
Xinyang Maojian (XYMJ) tea is one of the world's most popular green teas; the development of new sprouts directly affects the yield and quality of tea products, especially for XYMJ, which has hairy tips. Here, we used transcriptome and small RNA sequencing to identify mRNAs and miRNAs, respectively, involved in regulating leaf development in different plant tissues (bud, leaf, and stem). We identified a total of 381 conserved miRNAs. Given that no genomic data for XYMJ green tea are available, we compared the sequencing data for XYMJ green tea with genomic data from a closely related species (Tieguanyin) and the Camellia sinensis var. sinensis database; we identified a total of 506 and 485 novel miRNAs, respectively. We also identified 11 sequence-identical novel miRNAs in the tissues of XYMJ tea plants. Correlation analyses revealed 97 miRNA-mRNA pairs involved in leaf growth and development; the csn-miR319-2/csnTCP2 and miR159-csnMYB modules were found to be involved in leaf development in XYMJ green tea. Quantitative real-time PCR was used to validate the expression levels of the miRNAs and mRNAs. The miRNAs and target genes identified in this study might shed new light on the molecular mechanisms underlying the regulation of leaf development in tea plants.
Keywords: Xinyang Maojian; leaf development; miRNA; regulatory network; tea plant.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Dynamic Changes of Volatile Compounds during the Xinyang Maojian Green Tea Manufacturing at an Industrial Scale.Foods. 2022 Sep 2;11(17):2682. doi: 10.3390/foods11172682. Foods. 2022. PMID: 36076866 Free PMC article.
-
Genome-wide identification of conserved and novel microRNAs in one bud and two tender leaves of tea plant (Camellia sinensis) by small RNA sequencing, microarray-based hybridization and genome survey scaffold sequences.BMC Plant Biol. 2017 Nov 21;17(1):212. doi: 10.1186/s12870-017-1169-1. BMC Plant Biol. 2017. PMID: 29157210 Free PMC article.
-
Characterization of Key Odorants in Xinyang Maojian Green Tea and Their Changes During the Manufacturing Process.J Agric Food Chem. 2022 Jan 12;70(1):279-288. doi: 10.1021/acs.jafc.1c06473. Epub 2021 Dec 21. J Agric Food Chem. 2022. PMID: 34932338
-
Transcriptome-based variations effectively untangling the intraspecific relationships and selection signals in Xinyang Maojian tea population.Front Plant Sci. 2023 Feb 20;14:1114284. doi: 10.3389/fpls.2023.1114284. eCollection 2023. Front Plant Sci. 2023. PMID: 36890899 Free PMC article.
-
Green tea actions on miRNAs expression - An update.Chem Biol Interact. 2023 Jun 1;378:110465. doi: 10.1016/j.cbi.2023.110465. Epub 2023 Mar 31. Chem Biol Interact. 2023. PMID: 37004950 Review.
Cited by
-
Uncovering the key miRNA-target network of tea plants in resistance to sooty mold disease.BMC Plant Biol. 2025 Apr 9;25(1):446. doi: 10.1186/s12870-025-06509-7. BMC Plant Biol. 2025. PMID: 40200131 Free PMC article.
References
-
- Wei C., Yang H., Wang S., Zhao J., Liu C., Gao L., Xia E., Lu Y., Tai Y., She G., et al. Draft genome sequence of Camellia sinensis var. sinensis provides insights into the evolution of the tea genome and tea quality. Proc. Natl. Acad. Sci. USA. 2018;115:E4151–E4158. doi: 10.1073/pnas.1719622115. - DOI - PMC - PubMed
-
- Ming T., Bartholomew B. Theaceae. In: Wu Z., Raven P., Hong D., editors. Flora of China. Science Press and Missouri Botanical Garden; Beijing, China: St. Louis, MI, USA: 2007. pp. 367–412.
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

