GhDof1.7, a Dof Transcription Factor, Plays Positive Regulatory Role under Salinity Stress in Upland Cotton
- PMID: 37960096
- PMCID: PMC10649836
- DOI: 10.3390/plants12213740
GhDof1.7, a Dof Transcription Factor, Plays Positive Regulatory Role under Salinity Stress in Upland Cotton
Abstract
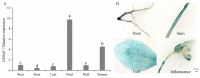
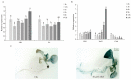
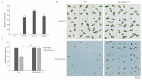
Salt stress is a major abiotic stressor that can severely limit plant growth, distribution, and crop yield. DNA-binding with one finger (Dof) is a plant-specific transcription factor that plays a crucial role in plant growth, development, and stress response. In this study, the function of a Dof transcription factor, GhDof1.7, was investigated in upland cotton. The GhDof1.7 gene has a coding sequence length of 759 base pairs, encoding 252 amino acids, and is mainly expressed in roots, stems, leaves, and inflorescences. Salt and abscisic acid (ABA) treatments significantly induced the expression of GhDof1.7. The presence of GhDof1.7 in Arabidopsis may have resulted in potential improvements in salt tolerance, as suggested by a decrease in H2O2 content and an increase in catalase (CAT) and superoxide dismutase (SOD) activities. The GhDof1.7 protein was found to interact with GhCAR4 (C2-domain ABA-related 4), and the silencing of either GhDof1.7 or GhCAR4 resulted in reduced salt tolerance in cotton plants. These findings demonstrate that GhDof1.7 plays a crucial role in improving the salt tolerance of upland cotton and provide insight into the regulation of abiotic stress response by Dof transcription factors.
Keywords: ABA; DNA-binding with one finger; H2O2; salt stress; upland cotton.
Conflict of interest statement
The authors declared that they have no conflict of interest to this work.
Figures


Similar articles
-
Overexpression of GhDof1 improved salt and cold tolerance and seed oil content in Gossypium hirsutum.J Plant Physiol. 2017 Nov;218:222-234. doi: 10.1016/j.jplph.2017.07.017. Epub 2017 Jul 28. J Plant Physiol. 2017. PMID: 28888163
-
Overexpression of GhABF3 increases cotton(Gossypium hirsutum L.) tolerance to salt and drought.BMC Plant Biol. 2022 Jun 29;22(1):313. doi: 10.1186/s12870-022-03705-7. BMC Plant Biol. 2022. PMID: 35768771 Free PMC article.
-
Temporal salt stress-induced transcriptome alterations and regulatory mechanisms revealed by PacBio long-reads RNA sequencing in Gossypium hirsutum.BMC Genomics. 2020 Nov 27;21(1):838. doi: 10.1186/s12864-020-07260-z. BMC Genomics. 2020. PMID: 33246403 Free PMC article.
-
ABP9, a maize bZIP transcription factor, enhances tolerance to salt and drought in transgenic cotton.Planta. 2017 Sep;246(3):453-469. doi: 10.1007/s00425-017-2704-x. Epub 2017 May 4. Planta. 2017. PMID: 28474114
-
The CCCH-Type Zinc-Finger Protein GhC3H20 Enhances Salt Stress Tolerance in Arabidopsis thaliana and Cotton through ABA Signal Transduction Pathway.Int J Mol Sci. 2023 Mar 6;24(5):5057. doi: 10.3390/ijms24055057. Int J Mol Sci. 2023. PMID: 36902489 Free PMC article.
Cited by
-
Identification of the Dof Gene Family in Quinoa and Its Potential Role in Regulating Flavonoid Synthesis Under Different Stress Conditions.Biology (Basel). 2025 Apr 20;14(4):446. doi: 10.3390/biology14040446. Biology (Basel). 2025. PMID: 40282311 Free PMC article.
-
Genome-Wide Identification and Expression Analysis of the LbDof Transcription Factor Family Genes in Lycium barbarum.Plants (Basel). 2025 May 22;14(11):1567. doi: 10.3390/plants14111567. Plants (Basel). 2025. PMID: 40508247 Free PMC article.
-
Genome-wide identification and molecular evolution of Dof gene family in Camellia oleifera.BMC Genomics. 2024 Jul 18;25(1):702. doi: 10.1186/s12864-024-10622-6. BMC Genomics. 2024. PMID: 39026173 Free PMC article.
-
Genome-wide identification and expression analysis of Dof gene family members in mulberry trees (Morus notabilis L.) under drought stress.BMC Genomics. 2025 Aug 12;26(1):744. doi: 10.1186/s12864-025-11843-z. BMC Genomics. 2025. PMID: 40797181 Free PMC article.
-
Comprehensive co-expression network reveals the fine-tuning of AsHSFA2c in balancing drought tolerance and growth in oat.Commun Biol. 2025 Mar 8;8(1):393. doi: 10.1038/s42003-025-07857-8. Commun Biol. 2025. PMID: 40057657 Free PMC article.
References
-
- Liu J., Guo W.Q., Shi D.C. Seed germination, seedling survival, and physiological response of sunflowers under saline and alkaline conditions. Photosynthetica. 2010;48:278–286.
-
- Mahajan S., Tuteja N. Cold, salinity and drought stresses: An overview. Arch. Biochem. Biophys. 2005;444:139–158. - PubMed
-
- Rehman S., Abbas G., Shahid M., Saqib M., Umer Farooq A.B., Hussain M., Murtaza B., Amjad M., Naeem M.A., Farooq A. Effect of salinity on cadmium tolerance, ionic homeostasis and oxidative stress responses in conocarpus exposed to cadmium stress: Implications for phytoremediation. Ecotoxicol. Environ. Saf. 2019;171:146–153. - PubMed
-
- Munns R. Comparative physiology of salt and water stress. Plant Cell Environ. 2002;25:239–250. - PubMed
-
- Gill S.S., Tuteja N. Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiol. Biochem. 2010;48:909–930. - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous

