This is a preprint.
Last in first out: SIV proviruses seeded later in infection are harbored in short-lived CD4+ T cells
- PMID: 37961482
- PMCID: PMC10635124
- DOI: 10.1101/2023.11.03.565539
Last in first out: SIV proviruses seeded later in infection are harbored in short-lived CD4+ T cells
Update in
-
SIV proviruses seeded later in infection are harbored in short-lived CD4+ T cells.Cell Rep. 2025 May 27;44(5):115663. doi: 10.1016/j.celrep.2025.115663. Epub 2025 May 5. Cell Rep. 2025. PMID: 40327506 Free PMC article.
Abstract
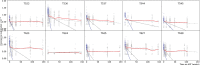
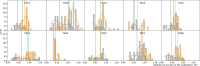
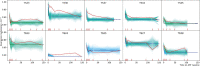
HIV can persist in a latent form as integrated DNA (provirus) in resting CD4+ T cells of infected individuals and as such is unaffected by antiretroviral therapy (ART). Despite being a major obstacle for eradication efforts, the genetic variation and timing of formation of this latent reservoir remains poorly understood. Previous studies on when virus is deposited in the latent reservoir have come to contradictory conclusions. To reexamine the genetic variation of HIV in CD4+ T cells during ART, we determined the divergence in envelope sequences collected from 10 SIV infected rhesus macaques. We found that the macaques displayed a biphasic decline of the viral divergence over time, where the first phase lasted for an average of 11.6 weeks (range 4-28 weeks). Motivated by recent observations that the HIV-infected CD4+ T cell population is composed of short- and long-lived subsets, we developed a model to study the divergence dynamics. We found that SIV in short-lived cells was on average more diverged, while long-lived cells harbored less diverged virus. This suggests that the long-lived cells harbor virus deposited starting earlier in infection and continuing throughout infection, while short-lived cells predominantly harbor more recent virus. As these cell populations decayed, the overall proviral divergence decline matched that observed in the empirical data. This model explains previous seemingly contradictory results on the timing of virus deposition into the latent reservoir, and should provide guidance for future eradication efforts.
Keywords: HIV; SIV; antiviral treatment; genetic divergence; latency.
Figures

References
-
- CDC centers for disease control, HIV and AIDS timeline (https://npin.cdc.gov/pages/hiv-and-aids-timeline) (2022) Accessed: 2022-08-24.
-
- WHO world health organization (https://www.who.int/data/gho/data/themes/hiv-aids) (2022) Accessed: 2022-08-24.
-
- Finzi D, et al. , Latent infection of CD4+ T cells provides a mechanism for lifelong persistence of HIV-1, even in patients on effective combination therapy. Nat. medicine 5, 512–517 (1999). - PubMed
-
- Wong JK, et al. , Recovery of replication-competent HIV despite prolonged suppression of plasma viremia. Science 278, 1291–1295 (1997). - PubMed
Publication types
Grants and funding
- R01 OD011095/OD/NIH HHS/United States
- R01 AI087520/AI/NIAID NIH HHS/United States
- P30 AI094189/AI/NIAID NIH HHS/United States
- UM1 AI164561/AI/NIAID NIH HHS/United States
- R01 AI149670/AI/NIAID NIH HHS/United States
- R01 AI028433/AI/NIAID NIH HHS/United States
- P01 AI169615/AI/NIAID NIH HHS/United States
- UM1 AI164556/AI/NIAID NIH HHS/United States
- DP5 OD031834/OD/NIH HHS/United States
- UM1 AI124377/AI/NIAID NIH HHS/United States
- U19 AI128751/AI/NIAID NIH HHS/United States
- UM1 AI164566/AI/NIAID NIH HHS/United States
LinkOut - more resources
Full Text Sources
Research Materials
