Chloroplast phosphate transporter CrPHT4-7 regulates phosphate homeostasis and photosynthesis in Chlamydomonas
- PMID: 37962583
- PMCID: PMC10904345
- DOI: 10.1093/plphys/kiad607
Chloroplast phosphate transporter CrPHT4-7 regulates phosphate homeostasis and photosynthesis in Chlamydomonas
Abstract
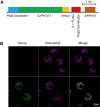
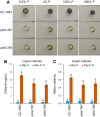
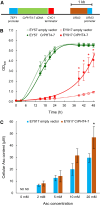
In eukaryotic cells, phosphorus is assimilated and utilized primarily as phosphate (Pi). Pi homeostasis is mediated by transporters that have not yet been adequately characterized in green algae. This study reports on PHOSPHATE TRANSPORTER 4-7 (CrPHT4-7) from Chlamydomonas reinhardtii, a member of the PHT4 transporter family, which exhibits remarkable similarity to AtPHT4;4 from Arabidopsis (Arabidopsis thaliana), a chloroplastic ascorbate transporter. Using fluorescent protein tagging, we show that CrPHT4-7 resides in the chloroplast envelope membrane. Crpht4-7 mutants, generated by the CRISPR/Cas12a-mediated single-strand templated repair, show retarded growth, especially in high light, reduced ATP level, strong ascorbate accumulation, and diminished non-photochemical quenching in high light. On the other hand, total cellular phosphorous content was unaffected, and the phenotype of the Crpht4-7 mutants could not be alleviated by ample Pi supply. CrPHT4-7-overexpressing lines exhibit enhanced biomass accumulation under high light conditions in comparison with the wild-type strain. Expressing CrPHT4-7 in a yeast (Saccharomyces cerevisiae) strain lacking Pi transporters substantially recovered its slow growth phenotype, demonstrating that CrPHT4-7 transports Pi. Even though CrPHT4-7 shows a high degree of similarity to AtPHT4;4, it does not display any substantial ascorbate transport activity in yeast or intact algal cells. Thus, the results demonstrate that CrPHT4-7 functions as a chloroplastic Pi transporter essential for maintaining Pi homeostasis and photosynthesis in C. reinhardtii.
© The Author(s) 2023. Published by Oxford University Press on behalf of American Society of Plant Biologists.
Conflict of interest statement
Conflict of interest statement. None declared.
Figures





Similar articles
-
The thylakoid- and pyrenoid-localized phosphate transporter PHT4-9 is essential for photosynthesis in Chlamydomonas.Plant Physiol. 2025 Apr 30;198(1):kiaf158. doi: 10.1093/plphys/kiaf158. Plant Physiol. 2025. PMID: 40273387 Free PMC article.
-
AtPHT4;4 is a chloroplast-localized ascorbate transporter in Arabidopsis.Nat Commun. 2015 Jan 5;6:5928. doi: 10.1038/ncomms6928. Nat Commun. 2015. PMID: 25557369 Free PMC article.
-
Genome-Wide Identification, Expression Profiling, and Evolution of Phosphate Transporter Gene Family in Green Algae.Front Genet. 2020 Oct 8;11:590947. doi: 10.3389/fgene.2020.590947. eCollection 2020. Front Genet. 2020. PMID: 33133172 Free PMC article.
-
Functional genomics of plant photosynthesis in the fast lane using Chlamydomonas reinhardtii.Trends Plant Sci. 2001 Aug;6(8):364-71. doi: 10.1016/s1360-1385(01)02018-0. Trends Plant Sci. 2001. PMID: 11495790 Review.
-
Chloroplast sulfate transport in green algae--genes, proteins and effects.Photosynth Res. 2005 Dec;86(3):299-307. doi: 10.1007/s11120-005-7382-z. Epub 2005 Nov 12. Photosynth Res. 2005. PMID: 16307303 Review.
Cited by
-
Transporting to the matrix: A pyrenoid-localized phosphate transporter required for optimal photoautotrophic growth in algae.Plant Physiol. 2025 May 30;198(2):kiaf232. doi: 10.1093/plphys/kiaf232. Plant Physiol. 2025. PMID: 40473241 Free PMC article. No abstract available.
-
The thylakoid- and pyrenoid-localized phosphate transporter PHT4-9 is essential for photosynthesis in Chlamydomonas.Plant Physiol. 2025 Apr 30;198(1):kiaf158. doi: 10.1093/plphys/kiaf158. Plant Physiol. 2025. PMID: 40273387 Free PMC article.
-
Effects of suppression of phosphate transporter 4;4 on CO2 assimilation in rice.J Plant Res. 2025 Jul;138(4):667-677. doi: 10.1007/s10265-025-01638-4. Epub 2025 Apr 18. J Plant Res. 2025. PMID: 40251403 Free PMC article.
-
Genome-wide association study reveals genetic basis and candidate genes for chlorophyll content of leaves in maize (Zea mays L.).PeerJ. 2024 Oct 7;12:e18278. doi: 10.7717/peerj.18278. eCollection 2024. PeerJ. 2024. PMID: 39391824 Free PMC article.
-
Genome-wide identification, expression analysis, and response to abiotic stress of the phosphate transporter gene family in cucumber (Cucumis sativus L.).BMC Plant Biol. 2025 May 21;25(1):677. doi: 10.1186/s12870-025-06720-6. BMC Plant Biol. 2025. PMID: 40399819 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

