Experimental and computational methods to highlight behavioural variations in TonB-dependent transporter expression in Pseudomonas aeruginosa versus siderophore concentration
- PMID: 37974013
- PMCID: PMC10654771
- DOI: 10.1038/s41598-023-46585-z
Experimental and computational methods to highlight behavioural variations in TonB-dependent transporter expression in Pseudomonas aeruginosa versus siderophore concentration
Abstract
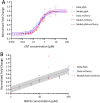
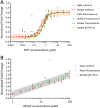
Iron is a key nutrient for bacterial growth. The source can be either heme or siderophore-Fe complexes. Siderophores are small molecules synthesized by bacteria to scavenge iron from the bacterial environment. The pathogen Pseudomonas aeruginosa can express at least 15 different iron uptake pathways and all but one involve a TonB-dependent transporter (TBDT) for the uptake of iron across the outer membrane. Little is known about how bacteria modulate and adapt the expression of their different iron import pathways according to their environment. Here, we have developed fluorescent reporters between the promoter region of genes encoding a TBDT and the fluorescent reporter mCherry. With these constructs, we can follow the expression of TBDTs under different growth conditions. Mathematical modelling of the data obtained showed the transcription and expression of the gene encoding the TBDT PfeA to have a sigmoidal shape, whereas it was logarithmic for the TBDT gene foxA. Maximum transcription for pfeA was reached in the presence of 3 µM enterobactin, the siderophore recognized by PfeA, whereas the maximum was not reached for foxA with 100 µM nocardamine, the siderophore of FoxA.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures




Similar articles
-
The role of FoxA, FiuA, and FpvB in iron acquisition via hydroxamate-type siderophores in Pseudomonas aeruginosa.Sci Rep. 2024 Aug 13;14(1):18795. doi: 10.1038/s41598-024-69152-6. Sci Rep. 2024. PMID: 39138320 Free PMC article.
-
The Pseudomonas aeruginosa pirA gene encodes a second receptor for ferrienterobactin and synthetic catecholate analogues.FEMS Microbiol Lett. 2005 May 15;246(2):167-74. doi: 10.1016/j.femsle.2005.04.010. FEMS Microbiol Lett. 2005. PMID: 15899402
-
Pseudomonas aeruginosa FpvB Is a High-Affinity Transporter for Xenosiderophores Ferrichrome and Ferrioxamine B.mBio. 2023 Feb 28;14(1):e0314922. doi: 10.1128/mbio.03149-22. Epub 2022 Dec 12. mBio. 2023. PMID: 36507834 Free PMC article.
-
Structure, function and binding selectivity and stereoselectivity of siderophore-iron outer membrane transporters.Curr Top Membr. 2012;69:37-66. doi: 10.1016/B978-0-12-394390-3.00002-1. Curr Top Membr. 2012. PMID: 23046646 Review.
-
Fate of ferrisiderophores after import across bacterial outer membranes: different iron release strategies are observed in the cytoplasm or periplasm depending on the siderophore pathways.Amino Acids. 2013 May;44(5):1267-77. doi: 10.1007/s00726-013-1468-2. Epub 2013 Feb 27. Amino Acids. 2013. PMID: 23443998 Review.
References
-
- Bullen, J. J., Rogers, H. J. & Griffiths, E. Role of iron in bacterial infection. In Curr. Top. Microbiol. Immunol. Vol. 80 (eds Arber, W. et al.) 1–35 (Springer, 1978). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

