SARS-CoV-2 variant-specific differences in inhibiting the effects of the PKR-activated integrated stress response
- PMID: 37979658
- PMCID: PMC10716588
- DOI: 10.1016/j.virusres.2023.199271
SARS-CoV-2 variant-specific differences in inhibiting the effects of the PKR-activated integrated stress response
Abstract
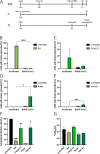
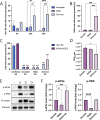
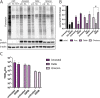
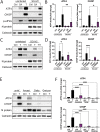
The integrated stress response (ISR) is a eukaryotic cell pathway that triggers translational arrest and the formation of stress granules (SGs) in response to various stress signals, including those caused by viral infections. The SARS-CoV-2 nucleocapsid protein has been shown to disrupt SGs, but SARS-CoV-2 interactions with other components of the pathway remains poorly characterized. Here, we show that SARS-CoV-2 infection triggers the ISR through activation of the eIF2α-kinase PKR while inhibiting a variety of downstream effects. In line with previous studies, SG formation was efficiently inhibited and the induced eIF2α phosphorylation only minimally contributed to the translational arrest observed in infected cells. Despite ISR activation and translational arrest, expression of the stress-responsive transcription factors ATF4 and CHOP was not induced in SARS-CoV-2 infected cells. Finally, we found variant-specific differences in the activation of the ISR between ancestral SARS-CoV-2 and the Delta and Omicron BA.1 variants in that Delta infection induced weaker PKR activation while Omicron infection induced higher levels of p-eIF2α, and greatly increased SG formation compared to the other variants. Our results suggest that different SARS-CoV-2 variants can affect normal cell functions differently, which can have an impact on pathogenesis and treatment strategies.
Keywords: Integrated stress response; Omicron; SARS-COV-2; SARS-COV-2 variants; Stress granules; Translational arrest.
Copyright © 2023 The Author(s). Published by Elsevier B.V. All rights reserved.
Conflict of interest statement
Declaration of Competing Interest The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures


Similar articles
-
The PERK/PKR-eIF2α Pathway Negatively Regulates Porcine Hemagglutinating Encephalomyelitis Virus Replication by Attenuating Global Protein Translation and Facilitating Stress Granule Formation.J Virol. 2022 Jan 12;96(1):e0169521. doi: 10.1128/JVI.01695-21. Epub 2021 Oct 13. J Virol. 2022. PMID: 34643429 Free PMC article.
-
Mouse Norovirus Infection Arrests Host Cell Translation Uncoupled from the Stress Granule-PKR-eIF2α Axis.mBio. 2019 Jun 18;10(3):e00960-19. doi: 10.1128/mBio.00960-19. mBio. 2019. PMID: 31213553 Free PMC article.
-
Peste Des Petits Ruminants Virus Nucleocapsid Protein Interacts with Protein Kinase R-Activating Protein and Induces Stress Granules To Promote Viral Replication.J Virol. 2023 Feb 28;97(2):e0171222. doi: 10.1128/jvi.01712-22. Epub 2023 Jan 18. J Virol. 2023. PMID: 36651745 Free PMC article.
-
Mammalian orthoreovirus escape from host translational shutoff correlates with stress granule disruption and is independent of eIF2alpha phosphorylation and PKR.J Virol. 2011 Sep;85(17):8798-810. doi: 10.1128/JVI.01831-10. Epub 2011 Jun 29. J Virol. 2011. PMID: 21715487 Free PMC article.
-
Hiding in Plain Sight: Formation and Function of Stress Granules During Microbial Infection of Mammalian Cells.Front Mol Biosci. 2021 Apr 29;8:647884. doi: 10.3389/fmolb.2021.647884. eCollection 2021. Front Mol Biosci. 2021. PMID: 33996904 Free PMC article. Review.
Cited by
-
A core network in the SARS-CoV-2 nucleocapsid NTD mediates structural integrity and selective RNA-binding.Nat Commun. 2024 Dec 9;15(1):10656. doi: 10.1038/s41467-024-55024-0. Nat Commun. 2024. PMID: 39653699 Free PMC article.
-
ATF4 Signaling in HIV-1 Infection: Viral Subversion of a Stress Response Transcription Factor.Biology (Basel). 2024 Feb 26;13(3):146. doi: 10.3390/biology13030146. Biology (Basel). 2024. PMID: 38534416 Free PMC article. Review.
-
Chasing Virus Replication and Infection: PAMP-PRR Interaction Drives Type I Interferon Production, Which in Turn Activates ISG Expression and ISGylation.Viruses. 2025 Apr 4;17(4):528. doi: 10.3390/v17040528. Viruses. 2025. PMID: 40284971 Free PMC article. Review.
References
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

