This is a preprint.
Single-cell transcriptomic and genomic changes in the aging human brain
- PMID: 37986960
- PMCID: PMC10659272
- DOI: 10.1101/2023.11.07.566050
Single-cell transcriptomic and genomic changes in the aging human brain
Update in
-
Single-cell transcriptomic and genomic changes in the ageing human brain.Nature. 2025 Oct;646(8085):657-666. doi: 10.1038/s41586-025-09435-8. Epub 2025 Sep 3. Nature. 2025. PMID: 40903571 Free PMC article.
Abstract
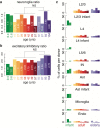
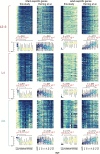
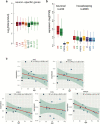
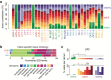
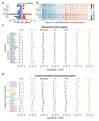
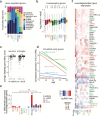
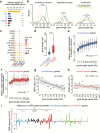
Aging brings dysregulation of various processes across organs and tissues, often stemming from stochastic damage to individual cells over time. Here, we used a combination of single-nucleus RNA-sequencing and single-cell whole-genome sequencing to identify transcriptomic and genomic changes in the prefrontal cortex of the human brain across life span, from infancy to centenarian. We identified infant-specific cell clusters enriched for the expression of neurodevelopmental genes, and a common down-regulation of cell-essential homeostatic genes that function in ribosomes, transport, and metabolism during aging across cell types. Conversely, expression of neuron-specific genes generally remains stable throughout life. We observed a decrease in specific DNA repair genes in aging, including genes implicated in generating brain somatic mutations as indicated by mutation signature analysis. Furthermore, we detected gene-length-specific somatic mutation rates that shape the transcriptomic landscape of the aged human brain. These findings elucidate critical aspects of human brain aging, shedding light on transcriptomic and genomics dynamics.
Figures









References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
