Integrative Proteogenomics for Differential Expression and Splicing Variation in a DM1 Mouse Model
- PMID: 37993104
- PMCID: PMC10770608
- DOI: 10.1016/j.mcpro.2023.100683
Integrative Proteogenomics for Differential Expression and Splicing Variation in a DM1 Mouse Model
Abstract
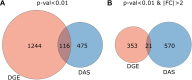
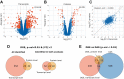
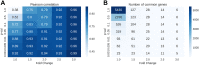
Dysregulated mRNA splicing is involved in the pathogenesis of many diseases including cancer, neurodegenerative diseases, and muscular dystrophies such as myotonic dystrophy type 1 (DM1). Comprehensive assessment of dysregulated splicing on the transcriptome and proteome level has been methodologically challenging, and thus investigations have often been targeting only few genes. Here, we performed a large-scale coordinated transcriptomic and proteomic analysis to characterize a DM1 mouse model (HSALR) in comparison to wild type. Our integrative proteogenomics approach comprised gene- and splicing-level assessments for mRNAs and proteins. It recapitulated many known instances of aberrant mRNA splicing in DM1 and identified new ones. It enabled the design and targeting of splicing-specific peptides and confirmed the translation of known instances of aberrantly spliced disease-related genes (e.g., Atp2a1, Bin1, Ryr1), complemented by novel findings (Flnc and Ywhae). Comparative analysis of large-scale mRNA and protein expression data showed quantitative agreement of differentially expressed genes and splicing patterns between disease and wild type. We hence propose this work as a suitable blueprint for a robust and scalable integrative proteogenomic strategy geared toward advancing our understanding of splicing-based disorders. With such a strategy, splicing-based biomarker candidates emerge as an attractive and accessible option, as they can be efficiently asserted on the mRNA and protein level in coordinated fashion.
Keywords: alternative splicing; myotonic dystrophy type 1 (DM1); proteogenomics.
Copyright © 2023 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Conflict of interest All authors are employees of Novartis and some hold Novartis stock.
Figures




References
-
- Bonnal S.C., López-Oreja I., Valcárcel J. Roles and mechanisms of alternative splicing in cancer — implications for care. Nat. Rev. Clin. Oncol. 2020;17:457–474. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

