ID1 expressing macrophages support cancer cell stemness and limit CD8+ T cell infiltration in colorectal cancer
- PMID: 37996458
- PMCID: PMC10667515
- DOI: 10.1038/s41467-023-43548-w
ID1 expressing macrophages support cancer cell stemness and limit CD8+ T cell infiltration in colorectal cancer
Abstract
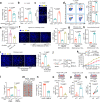
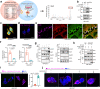
Elimination of cancer stem cells (CSCs) and reinvigoration of antitumor immunity remain unmet challenges for cancer therapy. Tumor-associated macrophages (TAMs) constitute the prominant population of immune cells in tumor tissues, contributing to the formation of CSC niches and a suppressive immune microenvironment. Here, we report that high expression of inhibitor of differentiation 1 (ID1) in TAMs correlates with poor outcome in patients with colorectal cancer (CRC). ID1 expressing macrophages maintain cancer stemness and impede CD8+ T cell infiltration. Mechanistically, ID1 interacts with STAT1 to induce its cytoplasmic distribution and inhibits STAT1-mediated SerpinB2 and CCL4 transcription, two secretory factors responsible for cancer stemness inhibition and CD8+ T cell recruitment. Reducing ID1 expression ameliorates CRC progression and enhances tumor sensitivity to immunotherapy and chemotherapy. Collectively, our study highlights the pivotal role of ID1 in controlling the protumor phenotype of TAMs and paves the way for therapeutic targeting of ID1 in CRC.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures





References
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

