Advances in Computational Approaches for Estimating Passive Permeability in Drug Discovery
- PMID: 37999336
- PMCID: PMC10673305
- DOI: 10.3390/membranes13110851
Advances in Computational Approaches for Estimating Passive Permeability in Drug Discovery
Abstract
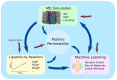
Passive permeation of cellular membranes is a key feature of many therapeutics. The relevance of passive permeability spans all biological systems as they all employ biomembranes for compartmentalization. A variety of computational techniques are currently utilized and under active development to facilitate the characterization of passive permeability. These methods include lipophilicity relations, molecular dynamics simulations, and machine learning, which vary in accuracy, complexity, and computational cost. This review briefly introduces the underlying theories, such as the prominent inhomogeneous solubility diffusion model, and covers a number of recent applications. Various machine-learning applications, which have demonstrated good potential for high-volume, data-driven permeability predictions, are also discussed. Due to the confluence of novel computational methods and next-generation exascale computers, we anticipate an exciting future for computationally driven permeability predictions.
Keywords: biomembrane; lipophilicity; machine learning; molecular dynamics; passive permeability.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources

