Assessing the Potential Role of Cats (Felis catus) as Generators of Relevant SARS-CoV-2 Lineages during the Pandemic
- PMID: 38003825
- PMCID: PMC10675002
- DOI: 10.3390/pathogens12111361
Assessing the Potential Role of Cats (Felis catus) as Generators of Relevant SARS-CoV-2 Lineages during the Pandemic
Abstract
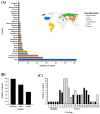
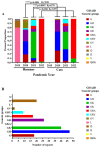
Several questions regarding the evolution of SARS-CoV-2 remain poorly elucidated. One of these questions is the possible evolutionary impact of SARS-CoV-2 after the infection in domestic animals. In this study, we aimed to evaluate the potential role of cats as generators of relevant SARS-CoV-2 lineages during the pandemic. A total of 105 full-length genome viral sequences obtained from naturally infected cats during the pandemic were evaluated by distinct evolutionary algorithms. Analyses were enhanced, including a set of highly related SARS-CoV-2 sequences recovered from human populations. Our results showed the apparent high susceptibility of cats to the infection SARS-CoV-2 compared with other animal species. Evolutionary analyses indicated that the phylogenomic characteristics displayed by cat populations were influenced by the dominance of specific SARS-CoV-2 genetic groups affecting human populations. However, disparate dN/dS rates at some genes between populations recovered from cats and humans suggested that infection in these two species may suggest a different evolutionary constraint for SARS-CoV-2. Interestingly, the branch selection analysis showed evidence of the potential role of natural selection in the emergence of five distinct cat lineages during the pandemic. Although these lineages were apparently irrelevant to public health during the pandemic, our results suggested that additional studies are needed to understand the role of other animal species in the evolution of SARS-CoV-2 during the pandemic.
Keywords: SARS-CoV-2; cats; evolution; phylogenomics; variants.
Conflict of interest statement
The authors declare no conflict of interest.
Figures





Similar articles
-
Tracking the Fate of Endogenous Retrovirus Segregation in Wild and Domestic Cats.J Virol. 2019 Nov 26;93(24):e01324-19. doi: 10.1128/JVI.01324-19. Print 2019 Dec 15. J Virol. 2019. PMID: 31534037 Free PMC article.
-
SARS-CoV-2 in domestic cats (Felis catus) in the northwest of Iran: Evidence for SARS-CoV-2 circulating between human and cats.Virus Res. 2022 Mar;310:198673. doi: 10.1016/j.virusres.2022.198673. Epub 2022 Jan 6. Virus Res. 2022. PMID: 34998863
-
Animals and SARS-CoV-2: Species susceptibility and viral transmission in experimental and natural conditions, and the potential implications for community transmission.Transbound Emerg Dis. 2021 Jul;68(4):1850-1867. doi: 10.1111/tbed.13885. Epub 2020 Nov 4. Transbound Emerg Dis. 2021. PMID: 33091230 Free PMC article. Review.
-
Severe SARS-CoV-2 Infection in a Cat with Hypertrophic Cardiomyopathy.Viruses. 2021 Jul 31;13(8):1510. doi: 10.3390/v13081510. Viruses. 2021. PMID: 34452375 Free PMC article.
-
[SARS-CoV-2 infections in cats, dogs, and other animal species: Findings on infection and data from Switzerland].Schweiz Arch Tierheilkd. 2021 Dec;163(12):821-835. doi: 10.17236/sat00329. Schweiz Arch Tierheilkd. 2021. PMID: 34881715 Review. German.
References
-
- Velazquez-Salinas L., Zarate S., Eberl S., Gladue D.P., Novella I., Borca M.V. Positive Selection of ORF1ab, ORF3a, and ORF8 Genes Drives the Early Evolutionary Trends of SARS-CoV-2 During the 2020 COVID-19 Pandemic. Front. Microbiol. 2020;11:550674. doi: 10.3389/fmicb.2020.550674. - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

