Molecular Characterization of Two Totiviruses from the Commensal Yeast Geotrichum candidum
- PMID: 38005831
- PMCID: PMC10674808
- DOI: 10.3390/v15112150
Molecular Characterization of Two Totiviruses from the Commensal Yeast Geotrichum candidum
Abstract
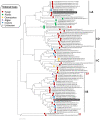
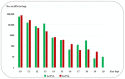
Mycoviruses can infect many of the major taxa of fungi including yeasts. Mycoviruses in the yeast fungus Geotrichum candidum are not well studied with only three G. candidum-associated viral species characterized to date, all of which belong to the Totiviridae genus Totivirus. In this study, we report the molecular characteristics of another two totiviruses co-infecting isolate Gc6 of G. candidum. The two totiviruses were tentatively named Geotrichum candidum totivirus 2 isolate Gc6 (GcTV2-Gc6) and Geotrichum candidum totivirus 4 isolate Gc6 (GcTV4-Gc6). Both viruses have the typical genome organization of totiviruses comprising two ORFs encoding capsid protein (CP) and RNA-dependent RNA polymerase (RdRp) at the N and C termini, respectively. The genomes of GcTV2-Gc6 and GcTV4-Gc6 are 4592 and 4530 bp long, respectively. Both viruses contain the-frameshifting elements and their proteins could be expressed as a single fusion protein. GcTV2-Gc6 is closely related to a totivirus isolated from the same host whereas GcTV4-Gc6 is related to insect-associated totiviruses. The phylogenetic analysis indicated that GcTV2-Gc6 and GcTV4-Gc6 belong to two different sister clades, I-A and I-B, respectively. It is interesting that all viruses identified from G. candidum belong to the genus Totivirus; however, this might be due to the lack of research reporting the characterization of mycoviruses from this fungal host. It is possible that the RNA interference (RNAi) mechanism cannot actively suppress totivirus accumulation in G. candidum Gc6.
Keywords: Geotrichum; RNA interference; Totivirus; dsRNA; high-throughput sequencing; mycoviruses; siRNA; usRNA.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



Similar articles
-
Identification of novel totiviruses from the ascomycetous fungus Geotrichum candidum.Arch Virol. 2022 Dec;167(12):2833-2838. doi: 10.1007/s00705-022-05611-7. Epub 2022 Oct 22. Arch Virol. 2022. PMID: 36271949
-
Sequence and phylogenetic analyses of novel totivirus-like double-stranded RNAs from field-collected powdery mildew fungi.Virus Res. 2016 Feb 2;213:353-364. doi: 10.1016/j.virusres.2015.11.015. Epub 2015 Dec 1. Virus Res. 2016. PMID: 26592174
-
Molecular characterization of three novel mycoviruses in the plant pathogenic fungus Exobasidium.Virus Res. 2022 Jan 2;307:198608. doi: 10.1016/j.virusres.2021.198608. Epub 2021 Nov 10. Virus Res. 2022. PMID: 34774616
-
Safety assessment of dairy microorganisms: Geotrichum candidum.Int J Food Microbiol. 2008 Sep 1;126(3):327-32. doi: 10.1016/j.ijfoodmicro.2007.08.021. Epub 2007 Aug 22. Int J Food Microbiol. 2008. PMID: 17869364 Review.
-
Characterization of Two Novel Toti-Like Viruses Co-infecting the Atlantic Blue Crab, Callinectes sapidus, in Its Northern Range of the United States.Front Microbiol. 2022 Mar 3;13:855750. doi: 10.3389/fmicb.2022.855750. eCollection 2022. Front Microbiol. 2022. PMID: 35369474 Free PMC article. Review.
Cited by
-
Identification and Genome Characterization of a Novel Virus within the Genus Totivirus from Chinese Bayberry (Myrica rubra).Viruses. 2024 Feb 12;16(2):283. doi: 10.3390/v16020283. Viruses. 2024. PMID: 38400058 Free PMC article.
-
Discovery of novel 'sugarcane totivirus 1' from Saccharum officinarum by high-throughput sequencing.Mol Biol Rep. 2025 Jan 15;52(1):122. doi: 10.1007/s11033-025-10221-y. Mol Biol Rep. 2025. PMID: 39812930
References
-
- Kim Y.-K., Kim T.-S., Shim H.-S., Park K.-S., Yeh W.-H., Hong S.-J., Shim C.-K., Kim J.-S., Park J.-H., Han E.-J., et al. First Report of Sour Rot on Post-harvest Oriental Melon, Tomato, Cucumber, Potato, Pumpkin and Carrot Caused by Geotrichum candidum. Res. Plant Dis. 2011;17:232–234. doi: 10.5423/RPD.2011.17.2.232. - DOI
-
- Aziza M., Couriol C., Amrane A., Boutrou R. Evidences for synergistic effects of Geotrichum candidum on Penicillium camembertii growing on cheese juice. Enzyme Microb. Technol. 2005;37:218–224. doi: 10.1016/j.enzmictec.2005.03.003. - DOI
-
- Boutrou R., Kerriou L., Gassi J.-Y. Contribution of Geotrichum candidum to the proteolysis of soft cheese. Int. Dairy J. 2006;16:775–783. doi: 10.1016/j.idairyj.2005.07.007. - DOI
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

