PRMT1 orchestrates with SAMTOR to govern mTORC1 methionine sensing via Arg-methylation of NPRL2
- PMID: 38006878
- PMCID: PMC11192564
- DOI: 10.1016/j.cmet.2023.11.001
PRMT1 orchestrates with SAMTOR to govern mTORC1 methionine sensing via Arg-methylation of NPRL2
Abstract
Methionine is an essential branch of diverse nutrient inputs that dictate mTORC1 activation. In the absence of methionine, SAMTOR binds to GATOR1 and inhibits mTORC1 signaling. However, how mTORC1 is activated upon methionine stimulation remains largely elusive. Here, we report that PRMT1 senses methionine/SAM by utilizing SAM as a cofactor for an enzymatic activity-based regulation of mTORC1 signaling. Under methionine-sufficient conditions, elevated cytosolic SAM releases SAMTOR from GATOR1, which confers the association of PRMT1 with GATOR1. Subsequently, SAM-loaded PRMT1 methylates NPRL2, the catalytic subunit of GATOR1, thereby suppressing its GAP activity and leading to mTORC1 activation. Notably, genetic or pharmacological inhibition of PRMT1 impedes hepatic methionine sensing by mTORC1 and improves insulin sensitivity in aged mice, establishing the role of PRMT1-mediated methionine sensing at physiological levels. Thus, PRMT1 coordinates with SAMTOR to form the methionine-sensing apparatus of mTORC1 signaling.
Keywords: GATOR1; NPRL2; PRMT1; arginine methylation; mTOR; methionine sensing; nutrient sensing.
Copyright © 2023 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests W.W. is a co-founder and consultant for ReKindle Therapeutics.
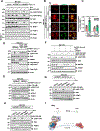
Figures





Similar articles
-
SAMTOR is an S-adenosylmethionine sensor for the mTORC1 pathway.Science. 2017 Nov 10;358(6364):813-818. doi: 10.1126/science.aao3265. Science. 2017. PMID: 29123071 Free PMC article.
-
Molecular mechanism of S-adenosylmethionine sensing by SAMTOR in mTORC1 signaling.Sci Adv. 2022 Jul;8(26):eabn3868. doi: 10.1126/sciadv.abn3868. Epub 2022 Jul 1. Sci Adv. 2022. PMID: 35776786 Free PMC article.
-
Arg-78 of Nprl2 catalyzes GATOR1-stimulated GTP hydrolysis by the Rag GTPases.J Biol Chem. 2019 Feb 22;294(8):2970-2975. doi: 10.1074/jbc.AC119.007382. Epub 2019 Jan 16. J Biol Chem. 2019. PMID: 30651352 Free PMC article.
-
Mechanism of Activation of Mechanistic Target of Rapamycin Complex 1 by Methionine.Front Cell Dev Biol. 2020 Aug 11;8:715. doi: 10.3389/fcell.2020.00715. eCollection 2020. Front Cell Dev Biol. 2020. PMID: 32850834 Free PMC article. Review.
-
Structures and Functions of the Human GATOR1 Complex.Subcell Biochem. 2024;104:269-294. doi: 10.1007/978-3-031-58843-3_12. Subcell Biochem. 2024. PMID: 38963491 Free PMC article. Review.
Cited by
-
The Molecular Basis of Amino Acids Sensing.Adv Sci (Weinh). 2025 Jul;12(26):e2501889. doi: 10.1002/advs.202501889. Epub 2025 May 24. Adv Sci (Weinh). 2025. PMID: 40411419 Free PMC article. Review.
-
Protein Arginine Methyltransferase 1: A Multi-Purpose Player in the Development of Cancer and Metabolic Disease.Biomolecules. 2025 Jan 27;15(2):185. doi: 10.3390/biom15020185. Biomolecules. 2025. PMID: 40001488 Free PMC article. Review.
-
Dysfunctional VLDL metabolism in MASLD.NPJ Metab Health Dis. 2024;2(1):16. doi: 10.1038/s44324-024-00018-1. Epub 2024 Jul 22. NPJ Metab Health Dis. 2024. PMID: 39049993 Free PMC article. Review.
-
mTORC1, the maestro of cell metabolism and growth.Genes Dev. 2025 Jan 7;39(1-2):109-131. doi: 10.1101/gad.352084.124. Genes Dev. 2025. PMID: 39572234 Free PMC article. Review.
-
Amino acids shape the metabolic and immunologic landscape in the tumor immune microenvironment: from molecular mechanisms to therapeutic strategies.Cancer Biol Med. 2025 Jul 24;22(7):726-46. doi: 10.20892/j.issn.2095-3941.2025.0115. Cancer Biol Med. 2025. PMID: 40708274 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

