Quantification of all 12 canonical ribonucleotides by real-time fluorogenic in vitro transcription
- PMID: 38008466
- PMCID: PMC10783517
- DOI: 10.1093/nar/gkad1091
Quantification of all 12 canonical ribonucleotides by real-time fluorogenic in vitro transcription
Abstract
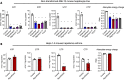
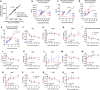
Enzymatic methods to quantify deoxyribonucleoside triphosphates have existed for decades. In contrast, no general enzymatic method to quantify ribonucleoside triphosphates (rNTPs), which drive almost all cellular processes and serve as precursors of RNA, exists to date. ATP can be measured with an enzymatic luminometric method employing firefly luciferase, but the quantification of other ribonucleoside mono-, di-, and triphosphates is still a challenge for a non-specialized laboratory and practically impossible without chromatography equipment. To allow feasible quantification of ribonucleoside phosphates in any laboratory with typical molecular biology and biochemistry tools, we developed a robust microplate assay based on real-time detection of the Broccoli RNA aptamer during in vitro transcription. The assay employs the bacteriophage T7 and SP6 RNA polymerases, two oligonucleotide templates encoding the 49-nucleotide Broccoli aptamer, and a high-affinity fluorogenic aptamer-binding dye to quantify each of the four canonical rNTPs. The inclusion of nucleoside mono- and diphosphate kinases in the assay reactions enabled the quantification of the mono- and diphosphate counterparts. The assay is inherently specific and tolerates concentrated tissue and cell extracts. In summary, we describe the first chromatography-free method to quantify ATP, ADP, AMP, GTP, GDP, GMP, UTP, UDP, UMP, CTP, CDP and CMP in biological samples.
© The Author(s) 2023. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures


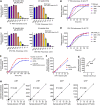

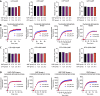
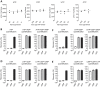
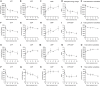
Similar articles
-
Discrimination of Ribonucleoside Mono-, Di-, and Triphosphates Using an Engineered Nanopore.ACS Nano. 2022 Dec 27;16(12):21356-21365. doi: 10.1021/acsnano.2c09662. Epub 2022 Dec 7. ACS Nano. 2022. PMID: 36475606
-
Formation of ribonucleotide 2',3'-cyclic carbonates during conversion of ribonucleoside 5'-phosphates to diphosphates and triphosphates by the phosphorimidazolidate procedure.Nucleic Acids Res. 1977 Aug;4(8):2843-53. doi: 10.1093/nar/4.8.2843. Nucleic Acids Res. 1977. PMID: 909793 Free PMC article.
-
Isocratic HPLC analysis for the simultaneous determination of dNTPs, rNTPs and ADP in biological samples.Nucleic Acids Res. 2022 Feb 22;50(3):e18. doi: 10.1093/nar/gkab1117. Nucleic Acids Res. 2022. PMID: 34850106 Free PMC article.
-
Chemoenzymatic preparation of nucleoside triphosphates.Curr Protoc Nucleic Acid Chem. 2004 May;Chapter 13:Unit 13.2. doi: 10.1002/0471142700.nc1302s16. Curr Protoc Nucleic Acid Chem. 2004. PMID: 18428922 Review.
-
Enzymatic recognition of 2'-modified ribonucleoside 5'-triphosphates: towards the evolution of versatile aptamers.Chembiochem. 2012 Jan 2;13(1):19-25. doi: 10.1002/cbic.201100648. Epub 2011 Dec 12. Chembiochem. 2012. PMID: 22162282 Review.
Cited by
-
Broccoli aptamer allows quantitative transcription regulation studies in vitro.PLoS One. 2024 Jun 13;19(6):e0304677. doi: 10.1371/journal.pone.0304677. eCollection 2024. PLoS One. 2024. PMID: 38870160 Free PMC article.
-
Activating AMPK improves pathological phenotypes due to mtDNA depletion.FEBS J. 2025 May;292(9):2359-2380. doi: 10.1111/febs.70006. Epub 2025 Feb 7. FEBS J. 2025. PMID: 39918244 Free PMC article.
-
Glucose limitation protects cancer cells from apoptosis induced by pyrimidine restriction and replication inhibition.Nat Metab. 2024 Dec;6(12):2338-2353. doi: 10.1038/s42255-024-01166-w. Epub 2024 Nov 26. Nat Metab. 2024. PMID: 39592843
References
-
- Traut T.W. Physiological concentrations of purines and pyrimidines. Mol. Cell. Biochem. 1994; 140:1–22. - PubMed
-
- Zhu B., Wei H., Wang Q., Li F., Dai J., Yan C., Cheng Y.. A simultaneously quantitative method to profiling twenty endogenous nucleosides and nucleotides in cancer cells using UHPLC-MS/MS. Talanta. 2018; 179:615–623. - PubMed
-
- Zborníková E., Knejzlík Z., Hauryliuk V., Krásný L., Rejman D. Analysis of nucleotide pools in bacteria using HPLC-MS in HILIC mode. Talanta. 2019; 205:120161. - PubMed
-
- Kong Z., Jia S., Chabes A.L., Appelblad P., Lundmark R., Moritz T., Chabes A.. Simultaneous determination of ribonucleoside and deoxyribonucleoside triphosphates in biological samples by hydrophilic interaction liquid chromatography coupled with tandem mass spectrometry. Nucleic Acids Res. 2018; 46:e66. - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

