Upregulated expression of miR-4443 and miR-4488 in drug resistant melanomas promotes migratory and invasive phenotypes through downregulation of intermediate filament nestin
- PMID: 38008717
- PMCID: PMC10680267
- DOI: 10.1186/s13046-023-02878-9
Upregulated expression of miR-4443 and miR-4488 in drug resistant melanomas promotes migratory and invasive phenotypes through downregulation of intermediate filament nestin
Abstract
Background: BRAF-mutant melanoma patients benefit from the combinatorial treatments with BRAF and MEK inhibitors. However, acquired drug resistance strongly limits the efficacy of these targeted therapies in time. Recently, many findings have underscored the involvement of microRNAs as main drivers of drug resistance. In this context, we previously identified a subset of oncomiRs strongly up-regulated in drug-resistant melanomas. In this work, we shed light on the molecular role of two as yet poorly characterized oncomiRs, miR-4443 and miR-4488.
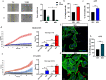
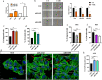
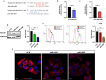
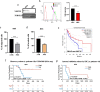
Methods: Invasion and migration have been determined by wound healing, transwell migration/invasion assays and Real Time Cell Analysis (RTCA) technology. miR-4488 and miR-4443 have been measured by qRT-PCR. Nestin levels have been tested by western blot, confocal immunofluorescence, immunohistochemical and flow cytometry analyses.
Results: We demonstrate that the two oncomiRs are responsible for the enhanced migratory and invasive phenotypes, that are a hallmark of drug resistant melanoma cells. Moreover, miR-4443 and miR-4488 promote an aberrant cytoskeletal reorganization witnessed by the increased number of stress fibers and cellular protrusions-like cancer cell invadopodia. Mechanistically, we identified the intermediate filament nestin as a molecular target of both oncomiRs. Finally, we have shown that nestin levels are able to predict response to treatments in melanoma patients.
Conclusions: Altogether these findings have profound translational implications in the attempt i) to develop miRNA-targeting therapies to mitigate the metastatic phenotypes of BRAF-mutant melanomas and ii) to identify novel biomarkers able to guide clinical decisions.
Keywords: Invasion; Metastatic melanoma; MicroRNA; Migration; Targeted therapy.
© 2023. The Author(s).
Conflict of interest statement
All the authors declare no competing interests with the exception of P.A.A. He has/had a consultant/advisory role for Bristol Myers Squibb, Roche-Genentech, Merck Sharp & Dohme, Novartis, Merck Serono, Pierre-Fabre, AstraZeneca, Sun Pharma, Sanofi, Idera, Sandoz, Immunocore, 4SC, Italfarmaco, Nektar, Boehringer-Ingelheim, Eisai, Regeneron, DaiichiSankyo, Pfizer, Oncosec, Nouscom, Lunaphore, Seagen, iTeos, Medicenna, Bio-Al Health, ValoTX, Replimmune. He also received research funding from Bristol Myers Squibb, Roche-Genentech, Pfizer, Sanofi. Travel support by Pfizer. The funders of this study had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Figures

Similar articles
-
Quantitative Proteomics Links the Intermediate Filament Nestin to Resistance to Targeted BRAF Inhibition in Melanoma Cells.Mol Cell Proteomics. 2019 Jun;18(6):1096-1109. doi: 10.1074/mcp.RA119.001302. Epub 2019 Mar 19. Mol Cell Proteomics. 2019. PMID: 30890564 Free PMC article.
-
miR-126-3p down-regulation contributes to dabrafenib acquired resistance in melanoma by up-regulating ADAM9 and VEGF-A.J Exp Clin Cancer Res. 2019 Jun 21;38(1):272. doi: 10.1186/s13046-019-1238-4. J Exp Clin Cancer Res. 2019. PMID: 31227006 Free PMC article.
-
miR-579-3p controls melanoma progression and resistance to target therapy.Proc Natl Acad Sci U S A. 2016 Aug 23;113(34):E5005-13. doi: 10.1073/pnas.1607753113. Epub 2016 Aug 8. Proc Natl Acad Sci U S A. 2016. PMID: 27503895 Free PMC article.
-
Upregulation of MicroRNA-1246 Is Associated with BRAF Inhibitor Resistance in Melanoma Cells with Mutant BRAF.Cancer Res Treat. 2017 Oct;49(4):947-959. doi: 10.4143/crt.2016.280. Epub 2017 Jan 3. Cancer Res Treat. 2017. PMID: 28052651 Free PMC article.
-
miR-146a-5p impairs melanoma resistance to kinase inhibitors by targeting COX2 and regulating NFkB-mediated inflammatory mediators.Cell Commun Signal. 2020 Sep 23;18(1):156. doi: 10.1186/s12964-020-00601-1. Cell Commun Signal. 2020. PMID: 32967672 Free PMC article.
Cited by
-
The MITF/mir-579-3p regulatory axis dictates BRAF-mutated melanoma cell fate in response to MAPK inhibitors.Cell Death Dis. 2024 Mar 12;15(3):208. doi: 10.1038/s41419-024-06580-2. Cell Death Dis. 2024. PMID: 38472212 Free PMC article.
-
Unveiling the Dynamic Interplay between Cancer Stem Cells and the Tumor Microenvironment in Melanoma: Implications for Novel Therapeutic Strategies.Cancers (Basel). 2024 Aug 16;16(16):2861. doi: 10.3390/cancers16162861. Cancers (Basel). 2024. PMID: 39199632 Free PMC article. Review.
-
Circulating MicroRNAs: functional biomarkers for melanoma prognosis and treatment.Mol Cancer. 2025 Mar 28;24(1):99. doi: 10.1186/s12943-025-02298-7. Mol Cancer. 2025. PMID: 40156012 Free PMC article. Review.
-
Development of an innovative duplex digital PCR assay for circulating MiRNA ratio quantification in metastatic melanoma.J Transl Med. 2025 Aug 13;23(1):904. doi: 10.1186/s12967-025-06941-1. J Transl Med. 2025. PMID: 40804405 Free PMC article.
-
Nestin in multiple myeloma: emerging insights into a potential therapeutic target.Front Oncol. 2025 Jul 29;15:1596928. doi: 10.3389/fonc.2025.1596928. eCollection 2025. Front Oncol. 2025. PMID: 40799240 Free PMC article. Review.
References
-
- Rastrelli M, Tropea S, Rossi CR, Alaibac M. Melanoma: epidemiology, risk factors, pathogenesis, diagnosis and classification. In Vivo. 2014;28(6):1005–11. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

