Draft genome sequencing of halotolerant bacterium Salinicola sp. DM10 unravels plant growth-promoting potentials
- PMID: 38009164
- PMCID: PMC10667196
- DOI: 10.1007/s13205-023-03833-3
Draft genome sequencing of halotolerant bacterium Salinicola sp. DM10 unravels plant growth-promoting potentials
Abstract
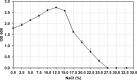
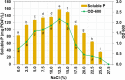
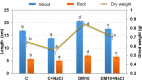
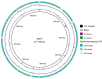
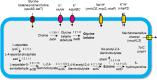
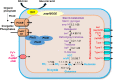
In this study, strain DM10 was isolated from mangrove roots and characterized as a halotolerant plant growth-promoting bacterium. Strain DM10 exhibited the ability to solubilize phosphate, produce siderophore, show 1-aminocyclopropane-1-carboxylic acid deaminase activity, and hydrolyze starch. The rice plants subjected to a treatment of NaCl (200 mM) and inoculated with strain DM10 showed an improvement in the shoot length, root length, and dried weight, when compared to those exposed solely to saline treatment. The comprehensive genome sequencing of strain DM10 revealed a genome spanning of 4,171,745 bp, harboring 3626 protein coding sequences. Within its genome, strain DM10 possesses genes responsible for both salt-in and salt-out strategies, indicative of a robust genetic adaptation aimed at fostering salt tolerance. Additionally, the genome encodes genes involved in phosphate solubilization, such as the synthesis of gluconic acid, high-affinity phosphate transport systems, and alkaline phosphatase. In the genome of DM10, we identified the acdS gene, responsible for encoding 1-aminocyclopropane-1-carboxylate deaminase, as well as the amy1A gene, which encodes α-amylase. Furthermore, the genome of DM10 contains sequences associated with the iron (3+)-hydroxamate and iron uptake clusters, responsible for siderophore production. Such data provide a deep understanding of the mechanism employed by strain DM10 to combat osmotic and salinity stress, facilitate plant growth, and elucidate its molecular-level behaviors.
Keywords: DM10; Genome sequencing; Halotolerant bacterium; Plant growth-promoting; Salinicola.
© King Abdulaziz City for Science and Technology 2023.
Conflict of interest statement
Conflict of interestThe authors declare no conflict of interest.
Figures


Similar articles
-
Mitigation of Salt Stress in Rice by the Halotolerant Plant Growth-Promoting Bacterium Enterobacter asburiae D2.J Xenobiot. 2024 Mar 1;14(1):333-349. doi: 10.3390/jox14010021. J Xenobiot. 2024. PMID: 38535496 Free PMC article.
-
Evaluation of Pseudomonas sp. for its multifarious plant growth promoting potential and its ability to alleviate biotic and abiotic stress in tomato (Solanum lycopersicum) plants.Sci Rep. 2020 Dec 1;10(1):20951. doi: 10.1038/s41598-020-77850-0. Sci Rep. 2020. PMID: 33262413 Free PMC article.
-
Achromobacter sp. FB-14 harboring ACC deaminase activity augmented rice growth by upregulating the expression of stress-responsive CIPK genes under salinity stress.Braz J Microbiol. 2020 Jun;51(2):719-728. doi: 10.1007/s42770-019-00199-8. Epub 2019 Dec 9. Braz J Microbiol. 2020. PMID: 31820296 Free PMC article.
-
Halotolerant Enterobacter asburiae A103 isolated from the halophyte Salix linearistipularis: Genomic analysis and growth-promoting effects on Medicago sativa under alkali stress.Microbiol Res. 2024 Dec;289:127909. doi: 10.1016/j.micres.2024.127909. Epub 2024 Sep 19. Microbiol Res. 2024. PMID: 39305780
-
Salinity stress endurance of the plants with the aid of bacterial genes.Front Genet. 2023 Apr 17;14:1049608. doi: 10.3389/fgene.2023.1049608. eCollection 2023. Front Genet. 2023. PMID: 37139239 Free PMC article. Review.
References
Publication types
LinkOut - more resources
Full Text Sources
Miscellaneous

