This is a preprint.
Intestinal cDC1s provide IL-12 dependent and independent functions required for CD4+ T cell-mediated resistance to Cryptosporidium
- PMID: 38014026
- PMCID: PMC10680586
- DOI: 10.1101/2023.11.11.566669
Intestinal cDC1s provide IL-12 dependent and independent functions required for CD4+ T cell-mediated resistance to Cryptosporidium
Update in
-
Intestinal cDC1s provide cues required for CD4+ T cell-mediated resistance to Cryptosporidium.J Exp Med. 2024 Jul 1;221(7):e20232067. doi: 10.1084/jem.20232067. Epub 2024 Jun 3. J Exp Med. 2024. PMID: 38829369 Free PMC article.
Abstract
Cryptosporidium is an enteric pathogen that is a prominent cause of diarrheal disease. Control of this infection requires CD4+ T cells, though the processes that lead to T cell-mediated resistance have been difficult to assess. Here, Cryptosporidium parasites that express MHCII-restricted model antigens were generated to dissect the early events that influence CD4+ T cell priming and effector function. These studies highlight that parasite-specific CD4+ T cells are primed in the draining mesenteric lymph node (mesLN) and differentiate into Th1 cells in the gut, where they mediate IFN-γ-dependent control of the infection. Although type 1 conventional dendritic cells (cDC1s) were not required for initial priming of CD4+ T cells, cDC1s were required for CD4+ T cell expansion and gut homing. cDC1s were also a major source of IL-12 that was not required for priming but promoted full differentiation of CD4+ T cells and local production of IFN-γ. Together, these studies reveal distinct roles for cDC1s in shaping CD4+ T cell responses to enteric infection: first to drive early expansion in the mesLN and second to drive effector responses in the gut.
Conflict of interest statement
Competing Interests: J.A.G. is currently affiliated with Cell Press, but all experiments performed by her for these studies were done before she worked there. Therefore, the authors declare no competing interests.
Figures




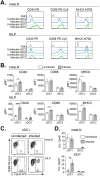


References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
