This is a preprint.
Modeling spatial evolution of multi-drug resistance under drug environmental gradients
- PMID: 38014279
- PMCID: PMC10680811
- DOI: 10.1101/2023.11.16.567447
Modeling spatial evolution of multi-drug resistance under drug environmental gradients
Update in
-
Modeling spatial evolution of multi-drug resistance under drug environmental gradients.PLoS Comput Biol. 2024 May 31;20(5):e1012098. doi: 10.1371/journal.pcbi.1012098. eCollection 2024 May. PLoS Comput Biol. 2024. PMID: 38820350 Free PMC article.
Abstract
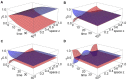
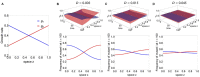
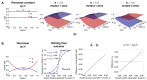
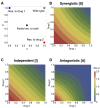
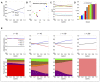
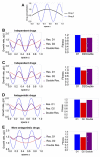
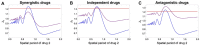
Multi-drug combinations to treat bacterial populations are at the forefront of approaches for infection control and prevention of antibiotic resistance. Although the evolution of antibiotic resistance has been theoretically studied with mathematical population dynamics models, extensions to spatial dynamics remain rare in the literature, including in particular spatial evolution of multi-drug resistance. In this study, we propose a reaction-diffusion system that describes the multi-drug evolution of bacteria, based on a rescaling approach (Gjini and Wood, 2021). We show how the resistance to drugs in space, and the consequent adaptation of growth rate is governed by a Price equation with diffusion. The covariance terms in this equation integrate features of drug interactions and collateral resistances or sensitivities to the drugs. We study spatial versions of the model where the distribution of drugs is homogeneous across space, and where the drugs vary environmentally in a piecewise-constant, linear and nonlinear manner. Applying concepts from perturbation theory and reaction-diffusion equations, we propose an analytical characterization of average mutant fitness in the spatial system based on the principal eigenvalue of our linear problem. This enables an accurate translation from drug spatial gradients and mutant antibiotic susceptibility traits, to the relative advantage of each mutant across the environment. Such a mathematical understanding allows to predict the precise outcomes of selection over space, ultimately from the fundamental balance between growth and movement traits, and their diversity in a population.
Keywords: environmental heterogeneity; infectious diseases; multi-drug antibiotic resistance; selection; spatial evolutionary dynamics.
Figures


Similar articles
-
Modeling spatial evolution of multi-drug resistance under drug environmental gradients.PLoS Comput Biol. 2024 May 31;20(5):e1012098. doi: 10.1371/journal.pcbi.1012098. eCollection 2024 May. PLoS Comput Biol. 2024. PMID: 38820350 Free PMC article.
-
Predictable properties of fitness landscapes induced by adaptational tradeoffs.Elife. 2020 May 19;9:e55155. doi: 10.7554/eLife.55155. Elife. 2020. PMID: 32423531 Free PMC article.
-
Price equation captures the role of drug interactions and collateral effects in the evolution of multidrug resistance.Elife. 2021 Jul 22;10:e64851. doi: 10.7554/eLife.64851. Elife. 2021. PMID: 34289932 Free PMC article.
-
Quantitative Clinical Imaging Methods for Monitoring Intratumoral Evolution.Methods Mol Biol. 2017;1513:61-81. doi: 10.1007/978-1-4939-6539-7_6. Methods Mol Biol. 2017. PMID: 27807831 Review.
-
Evolutionary Trajectories to Antibiotic Resistance.Annu Rev Microbiol. 2017 Sep 8;71:579-596. doi: 10.1146/annurev-micro-090816-093813. Epub 2017 Jul 11. Annu Rev Microbiol. 2017. PMID: 28697667 Review.
References
-
- Allen B., Lippner G., Chen Y.T., Fotouhi B., Momeni N., Yau S.T., Nowak M.A., 2017. Evolutionary dynamics on any population structure. Nature 544, 227–230. - PubMed
-
- Aronson D.G., Weinberger H.F., 1978. Multidimensional nonlinear diffusion arising in population genetics. Advances in Mathematics 30, 33–76.
-
- Asaduzzaman M., Rousham E., Unicomb L., Islam M.R., Amin M.B., Rahman M., Hossain M.I., Mahmud Z.H., Szegner M., Wood P., et al., 2022. Spatiotemporal distribution of antimicrobial resistant organisms in different water environments in urban and rural settings of bangladesh. Science of the Total Environment 831, 154890. - PubMed
-
- Atis S., Weinstein B.T., Murray A.W., Nelson D.R., 2019. Microbial range expansions on liquid substrates. Physical review X 9, 021058.
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
