Targeting PEG10 as a novel therapeutic approach to overcome CDK4/6 inhibitor resistance in breast cancer
- PMID: 38017459
- PMCID: PMC10683152
- DOI: 10.1186/s13046-023-02903-x
Targeting PEG10 as a novel therapeutic approach to overcome CDK4/6 inhibitor resistance in breast cancer
Abstract
Background: Breast cancer is the global leading cancer burden in women and the hormone receptor-positive (HR+) subtype is a major part of breast cancer. Though cyclin-dependent kinase 4 and 6 (CDK4/6) inhibitors are highly effective therapy for HR+ subtype, acquired resistance is inevitable in most cases. Herein, we investigated the paternally expressed gene 10 (PEG10)-associated mechanism of acquired resistance to CDK4/6 inhibitors.
Methods: Palbociclib-resistant cells were generated by exposing human HR+ breast cancer cell lines to palbociclib for 7-9 months. In vitro mechanistic study and in vivo xenograft assay were performed. For clinical relevance, public mRNA microarray data sets of early breast cancer were analyzed and PEG10 immunohistochemical staining was performed using pre-CDK4/6 inhibitor tumor samples.
Results: We observed that PEG10 was significantly upregulated in palbociclib-resistant cells. Ectopic overexpression of PEG10 in parental cells caused CDK4/6 inhibitor resistance and enhanced epithelial-mesenchymal transition (EMT). On the contrary, PEG10-targeting siRNA or antisense oligonucleotides (ASOs) combined with palbociclib synergistically inhibited proliferation of palbociclib-resistant cells and growth of palbociclib-resistant xenograft in mice and suppressed EMT as well. The mechanistic study confirmed that high PEG10 expression suppressed p21, a natural CDK inhibitor, and SIAH1, a post-translational degrader of ZEB1, augmenting CDK4/6 inhibitor resistance. Then PEG10 siRNA combined with palbociclib suppressed cell cycle progression and EMT via activating p21 and SIAH1, respectively. Consequently, combined PEG10 inhibition and palbociclib overcame CDK4/6 inhibitor resistance. Furthermore, high PEG10 expression was significantly associated with a shorter recurrence-free survival (RFS) based on public mRNA expression data. In pre-CDK4/6 inhibitor treatment tissues, PEG10 positivity by IHC also showed a trend toward a shorter progression-free survival (PFS) with CDK4/6 inhibitor. These results support clinical relevance of PEG10 as a therapeutic target.
Conclusions: We demonstrated a novel PEG10-associated mechanism of CDK4/6 inhibitor resistance. We propose PEG10 as a promising therapeutic target for overcoming PEG10-associated resistance to CDK4/6 inhibitors.
Keywords: ASO; CDK4/6; Drug resistance; HR+ breast cancer; PEG10.
© 2023. The Author(s).
Conflict of interest statement
The corresponding author received research funds from several pharmaceutical companies, including ImmunoMet Therapeutics, Hanmi Pharmaceutical Co., and Bixink Therapeutics. The authors declare no conflict of interest.
Figures


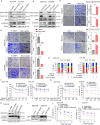



References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

