A machine learning and directed network optimization approach to uncover TP53 regulatory patterns
- PMID: 38047081
- PMCID: PMC10692668
- DOI: 10.1016/j.isci.2023.108291
A machine learning and directed network optimization approach to uncover TP53 regulatory patterns
Abstract
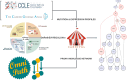
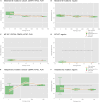
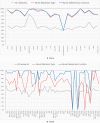
TP53, the Guardian of the Genome, is the most frequently mutated gene in human cancers and the functional characterization of its regulation is fundamental. To address this we employ two strategies: machine learning to predict the mutation status of TP53 from transcriptomic data, and directed regulatory networks to reconstruct the effect of mutations on the transcipt levels of TP53 targets. Using data from established databases (Cancer Cell Line Encyclopedia, The Cancer Genome Atlas), machine learning could predict the mutation status, but not resolve different mutations. On the contrary, directed network optimization allowed to infer the TP53 regulatory profile across: (1) mutations, (2) irradiation in lung cancer, and (3) hypoxia in breast cancer, and we could observe differential regulatory profiles dictated by (1) mutation type, (2) deleterious consequences of the mutation, (3) known hotspots, (4) protein changes, (5) stress condition (irradiation/hypoxia). This is an important first step toward using regulatory networks for the characterization of the functional consequences of mutations, and could be extended to other perturbations, with implications for drug design and precision medicine.
Keywords: Regulatory networks; TP53; cancer systems biology; causal inference; directed networks; machine learning; mutations; regulon; trascriptomics.
© 2024 The Authors.
Conflict of interest statement
J.S.R. reports funding from GSK, Pfizer, and Sanofi and fees/honoraria from Travere Therapeutics, Stadapharm, Astex, Pfizer, and Grunenthal.
Figures





References
-
- Yan W., Xue W., Chen J., Hu G. Biological networks for cancer candidate biomarkers discovery. Cancer Inf. 2016;15:1–7. https://pubmed.ncbi.nlm.nih.gov/27625573 - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

