Single-cell transcriptomics reveals variations in monocytes and Tregs between gout flare and remission
- PMID: 38063198
- PMCID: PMC10795830
- DOI: 10.1172/jci.insight.171417
Single-cell transcriptomics reveals variations in monocytes and Tregs between gout flare and remission
Erratum in
-
Single-cell transcriptomics reveals variations in monocytes and Tregs between gout flare and remission.JCI Insight. 2024 Feb 8;9(3):e179067. doi: 10.1172/jci.insight.179067. JCI Insight. 2024. PMID: 38329132 Free PMC article. No abstract available.
Abstract
Gout commonly manifests as a painful, self-limiting inflammatory arthritis. Nevertheless, the understanding of the inflammatory and immune responses underlying gout flares and remission remains ambiguous. Here, based on single-cell RNA-Seq and an independent validation cohort, we identified the potential mechanism of gout flare, which likely involves the upregulation of HLA-DQA1+ nonclassical monocytes and is related to antigen processing and presentation. Furthermore, Tregs also play an essential role in the suppressive capacity during gout remission. Cell communication analysis suggested the existence of altered crosstalk between monocytes and other T cell types, such as Tregs. Moreover, we observed the systemic upregulation of inflammatory and cytokine genes, primarily in classical monocytes, during gout flares. All monocyte subtypes showed increased arachidonic acid metabolic activity along with upregulation of prostaglandin-endoperoxide synthase 2 (PTGS2). We also detected a decrease in blood arachidonic acid and an increase in leukotriene B4 levels during gout flares. In summary, our study illustrates the distinctive immune cell responses and systemic inflammation patterns that characterize the transition from gout flares to remission, and it suggests that blood monocyte subtypes and Tregs are potential intervention targets for preventing recurrent gout attacks and progression.
Keywords: Immunology; Inflammation; Monocytes; T cells.
Figures
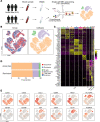





References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials

