A supernumerary synthetic chromosome in Komagataella phaffii as a repository for extraneous genetic material
- PMID: 38104077
- PMCID: PMC10724962
- DOI: 10.1186/s12934-023-02262-4
A supernumerary synthetic chromosome in Komagataella phaffii as a repository for extraneous genetic material
Abstract
Background: Komagataella phaffii (Pichia pastoris) is a methylotrophic commercially important non-conventional species of yeast that grows in a fermentor to exceptionally high densities on simple media and secretes recombinant proteins efficiently. Genetic engineering strategies are being explored in this organism to facilitate cost-effective biomanufacturing. Small, stable artificial chromosomes in K. phaffii could offer unique advantages by accommodating multiple integrations of extraneous genes and their promoters without accumulating perturbations of native chromosomes or exhausting the availability of selection markers.
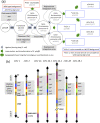
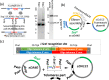
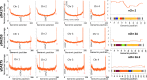
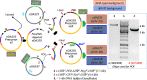
Results: Here, we describe a linear "nano"chromosome (of 15-25 kb) that, according to whole-genome sequencing, persists in K. phaffii over many generations with a copy number per cell of one, provided non-homologous end joining is compromised (by KU70-knockout). The nanochromosome includes a copy of the centromere from K. phaffii chromosome 3, a K. phaffii-derived autonomously replicating sequence on either side of the centromere, and a pair of K. phaffii-like telomeres. It contains, within its q arm, a landing zone in which genes of interest alternate with long (approx. 1-kb) non-coding DNA chosen to facilitate homologous recombination and serve as spacers. The landing zone can be extended along the nanochromosome, in an inch-worming mode of sequential gene integrations, accompanied by recycling of just two antibiotic-resistance markers. The nanochromosome was used to express PDI, a gene encoding protein disulfide isomerase. Co-expression with PDI allowed the production, from a genomically integrated gene, of secreted murine complement factor H, a plasma protein containing 40 disulfide bonds. As further proof-of-principle, we co-expressed, from a nanochromosome, both PDI and a gene for GFP-tagged human complement factor H under the control of PAOX1 and demonstrated that the secreted protein was active as a regulator of the complement system.
Conclusions: We have added K. phaffii to the list of organisms that can produce human proteins from genes carried on a stable, linear, artificial chromosome. We envisage using nanochromosomes as repositories for numerous extraneous genes, allowing intensive engineering of K. phaffii without compromising its genome or weakening the resulting strain.
Keywords: Artificial chromosome; Biomanufacture; Complement factor H; DNA assembly; Metabolic engineering; Pichia pastoris; Therapeutic proteins; Yeast.
© 2023. The Author(s).
Conflict of interest statement
None.
Figures



Similar articles
-
Komagataella phaffii (Pichia pastoris) as a Powerful Yeast Expression System for Biologics Production.Front Biosci (Elite Ed). 2024 Jun 12;16(2):19. doi: 10.31083/j.fbe1602019. Front Biosci (Elite Ed). 2024. PMID: 38939917 Review.
-
One-Step In Vivo Assembly of Multiple DNA Fragments and Genomic Integration in Komagataella phaffii.ACS Synth Biol. 2022 Feb 18;11(2):644-654. doi: 10.1021/acssynbio.1c00302. Epub 2022 Jan 30. ACS Synth Biol. 2022. PMID: 35094517
-
Comparative genomics and transcriptomics of Pichia pastoris.BMC Genomics. 2016 Aug 5;17:550. doi: 10.1186/s12864-016-2876-y. BMC Genomics. 2016. PMID: 27495311 Free PMC article.
-
Scalable protein production by Komagataella phaffii enabled by ARS plasmids and carbon source-based selection.Microb Cell Fact. 2024 Apr 20;23(1):116. doi: 10.1186/s12934-024-02368-3. Microb Cell Fact. 2024. PMID: 38643119 Free PMC article.
-
Industrial Production of Proteins with Pichia pastoris-Komagataella phaffii.Biomolecules. 2023 Feb 26;13(3):441. doi: 10.3390/biom13030441. Biomolecules. 2023. PMID: 36979376 Free PMC article. Review.
Cited by
-
Development of a Komagataella phaffii cell factory for sustainable production of ( +)-valencene.Microb Cell Fact. 2025 Jan 21;24(1):29. doi: 10.1186/s12934-025-02649-5. Microb Cell Fact. 2025. PMID: 39838465 Free PMC article.
-
Applications of the Methylotrophic Yeast Komagataella phaffii in the Context of Modern Biotechnology.J Fungi (Basel). 2024 Jun 6;10(6):411. doi: 10.3390/jof10060411. J Fungi (Basel). 2024. PMID: 38921397 Free PMC article. Review.
References
-
- Rebello S, Abraham A, Madhavan A, Sindhu R, Binod P, Karthika Bahuleyan A, Aneesh EM, Pandey A. Non-conventional yeast cell factories for sustainable bioprocesses. FEMS Microbiol Lett 2018, 365. - PubMed
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

