Identification of TFR2 as a novel ferroptosis‑related gene that serves an important role in prognosis and progression of triple‑negative breast cancer
- PMID: 38106522
- PMCID: PMC10722555
- DOI: 10.3892/ol.2023.14176
Identification of TFR2 as a novel ferroptosis‑related gene that serves an important role in prognosis and progression of triple‑negative breast cancer
Abstract
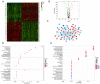
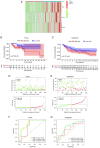
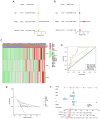
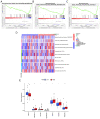
Effective targeted therapeutic strategies for triple-negative breast cancer (TNBC), the most malignant subtype of breast cancer, are currently lacking. Ferroptosis has been reported to be associated with the onset and advancement of various cancer types, including TNBC. However, there are limited studies on the correlation between TNBC and ferroptosis-related genes. In addition, the potential biomarkers of ferroptosis in TNBC need further investigation. The present study aimed to assess the prognostic role of a novel ferroptosis-related gene signature in the context of TNBC. The signature was established utilizing The Cancer Genome Atlas dataset. This three-gene model [transferrin receptor 2 (TFR2), regulator of G protein signaling 4 and zinc finger protein 36] was developed utilizing least absolute shrinkage and selection operator regression analysis and demonstrated satisfactory predictive performance in TNBC. The area under the curve values of the receiver operating characteristic curves in this model concerning the 1-, 2- and 3-year survival prediction were 0.721, 0.840 and 0.856, respectively. The predictive performance of the model was verified using the TNBC dataset GSE25307. Gene set enrichment analysis (GSEA) demonstrated the enrichment of genes in the low-risk group in a number of important metabolic pathways. Single-sample GSEA demonstrated a variation in the expression levels of immune checkpoint molecules between the high- and low-risk groups. The inhibitory impact of TFR2 knockdown on the proliferative capacity of TNBC cells was verified through in vitro experiments. The data also demonstrated that TFR2 knockdown facilitated the ferroptosis of TNBC cells. Additional assessments indicated that the effects of TFR2 knockdown were partially reversed upon treatment with the ferroptosis inhibitor ferrostatin-1. In conclusion, in the present study, a novel and accurate ferroptosis-related predictive signature was established for TNBC with potential future clinical applications. To the best of our knowledge, the present study is the first to report that TFR2 regulated ferroptosis in TNBC cells in vitro.
Keywords: ferroptosis; gene signature; prognosis; transferrin receptor 2; triple-negative breast cancer.
Copyright: © Yang et al.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures

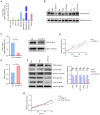
References
-
- Yu B, Luo F, Sun B, Liu W, Shi Q, Cheng SY, Chen C, Chen G, Li Y, Feng H. KAT6A Acetylation of SMAD3 regulates Myeloid-derived suppressor cell recruitment, metastasis, and immunotherapy in Triple-Negative breast cancer. Adv Sci (Weinh) 2021;8:e2100014. doi: 10.1002/advs.202100014. - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
