Metabolic engineering of high-salinity-induced biosynthesis of γ-aminobutyric acid improves salt-stress tolerance in a glutamic acid-overproducing mutant of an ectoine-deficient Halomonas elongata
- PMID: 38112419
- PMCID: PMC10807429
- DOI: 10.1128/aem.01905-23
Metabolic engineering of high-salinity-induced biosynthesis of γ-aminobutyric acid improves salt-stress tolerance in a glutamic acid-overproducing mutant of an ectoine-deficient Halomonas elongata
Abstract
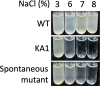
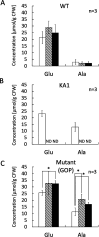
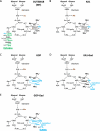
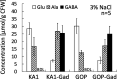
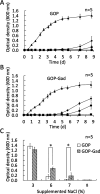
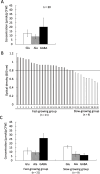
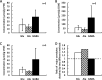
A moderately halophilic eubacterium, Halomonas elongata, has been used as cell factory to produce fine chemical 1,4,5,6-tetrahydro-2-methyl-4-pyrimidinecarboxylic acid (ectoine), which functions as a major osmolyte protecting the cells from high-salinity stress. To explore the possibility of using H. elongata to biosynthesize other valuable osmolytes, an ectoine-deficient salt-sensitive H. elongata deletion mutant strain KA1 (ΔectABC), which only grows well in minimal medium containing up to 3% NaCl, was subjected to an adaptive mutagenesis screening in search of mutants with restored salt tolerance. Consequently, we obtained a mutant, which tolerates 6% NaCl in minimal medium by overproducing L-glutamic acid (Glu). However, this Glu-overproducing (GOP) strain has a lower tolerance level than the wild-type H. elongata, possibly because the acidity of Glu interferes with the pH homeostasis of the cell and hinders its own cellular accumulation. Enzymatic decarboxylation of Glu to γ-aminobutyric acid (GABA) by a Glu decarboxylase (GAD) could restore cellular pH homeostasis; therefore, we introduced an engineered salt-inducible HopgadBmut gene, which encodes a wide pH-range GAD mutant, into the genome of the H. elongata GOP strain. We found that the resulting H. elongata GOP-Gad strain exhibits higher salt tolerance than the GOP strain by accumulating high concentration of GABA as an osmolyte in the cell (176.94 µmol/g cell dry weight in minimal medium containing 7% NaCl). With H. elongata OUT30018 genetic background, H. elongata GOP-Gad strain can utilize biomass-derived carbon and nitrogen compounds as its sole carbon and nitrogen sources, making it a good candidate for the development of GABA-producing cell factories.IMPORTANCEWhile the wild-type moderately halophilic H. elongata can synthesize ectoine as a high-value osmolyte via the aspartic acid metabolic pathway, a mutant H. elongata GOP strain identified in this work opens doors for the biosynthesis of alternative valuable osmolytes via glutamic acid metabolic pathway. Further metabolic engineering to install a GAD system into the H. elongata GOP strain successfully created a H. elongata GOP-Gad strain, which acquired higher tolerance to salt stress by accumulating GABA as a major osmolyte. With the ability to assimilate biomass-derived carbon and nitrogen sources and thrive in high-salinity environment, the H. elongata GOP-Gad strain can be used in the development of sustainable GABA-producing cell factories.
Keywords: Halomonas elongata; L-glutamic acid; compatible osmolyte; metabolic engineering; γ-aminobutyric acid.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



Similar articles
-
Enhanced accumulation of γ-aminobutyric acid by deletion of aminotransferase genes involved in γ-aminobutyric acid catabolism in engineered Halomonas elongata.Appl Environ Microbiol. 2024 Sep 18;90(9):e0073424. doi: 10.1128/aem.00734-24. Epub 2024 Aug 12. Appl Environ Microbiol. 2024. PMID: 39133003 Free PMC article.
-
Metabolic pathway engineering of high-salinity-induced overproduction of L-proline improves high-salinity stress tolerance of an ectoine-deficient Halomonas elongata.Appl Environ Microbiol. 2024 Sep 18;90(9):e0119524. doi: 10.1128/aem.01195-24. Epub 2024 Aug 19. Appl Environ Microbiol. 2024. PMID: 39158316 Free PMC article.
-
Expression of an engineered salt-inducible proline biosynthetic operon in a glutamic acid over-producing mutant, Halomonas elongata GOP, confers increased proline yield due to enhanced growth under high-salinity conditions.Biosci Biotechnol Biochem. 2024 Sep 20;88(10):1233-1241. doi: 10.1093/bbb/zbae102. Biosci Biotechnol Biochem. 2024. PMID: 39003245
-
Microbial production of ectoine and hydroxyectoine as high-value chemicals.Microb Cell Fact. 2021 Mar 26;20(1):76. doi: 10.1186/s12934-021-01567-6. Microb Cell Fact. 2021. PMID: 33771157 Free PMC article. Review.
-
[Advances in ectoine biosynthesis and biochemical characteristics of key enzymes].Sheng Wu Gong Cheng Xue Bao. 2024 Jun 25;40(6):1620-1643. doi: 10.13345/j.cjb.230640. Sheng Wu Gong Cheng Xue Bao. 2024. PMID: 38914483 Review. Chinese.
Cited by
-
Enhanced accumulation of γ-aminobutyric acid by deletion of aminotransferase genes involved in γ-aminobutyric acid catabolism in engineered Halomonas elongata.Appl Environ Microbiol. 2024 Sep 18;90(9):e0073424. doi: 10.1128/aem.00734-24. Epub 2024 Aug 12. Appl Environ Microbiol. 2024. PMID: 39133003 Free PMC article.
-
Hypersaline organic wastewater treatment: Biotechnological advances and engineering challenges.Environ Sci Ecotechnol. 2025 Feb 18;24:100542. doi: 10.1016/j.ese.2025.100542. eCollection 2025 Mar. Environ Sci Ecotechnol. 2025. PMID: 40083747 Free PMC article. Review.
-
Metabolic pathway engineering of high-salinity-induced overproduction of L-proline improves high-salinity stress tolerance of an ectoine-deficient Halomonas elongata.Appl Environ Microbiol. 2024 Sep 18;90(9):e0119524. doi: 10.1128/aem.01195-24. Epub 2024 Aug 19. Appl Environ Microbiol. 2024. PMID: 39158316 Free PMC article.
References
-
- Vreeland RH, Litchfield CD, Martin EL, Elliot E. 1980. Halomonas elongata, a new genus and species of extremely salt-tolerant bacteria. Int J Syst Evol Microbiol 30:485–495. doi:10.1099/00207713-30-2-485 - DOI
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

