Tolerant and Rapid Endochondral Bone Regeneration Using Framework-Enhanced 3D Biomineralized Matrix Hydrogels
- PMID: 38127989
- PMCID: PMC10916654
- DOI: 10.1002/advs.202305580
Tolerant and Rapid Endochondral Bone Regeneration Using Framework-Enhanced 3D Biomineralized Matrix Hydrogels
Abstract
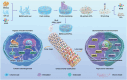
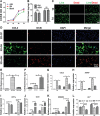
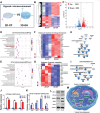
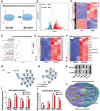
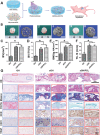
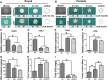
Tissue-engineered bone has emerged as a promising alternative for bone defect repair due to the advantages of regenerative bone healing and physiological functional reconstruction. However, there is very limited breakthrough in achieving favorable bone regeneration due to the harsh osteogenic microenvironment after bone injury, especially the avascular and hypoxic conditions. Inspired by the bone developmental mode of endochondral ossification, a novel strategy is proposed for tolerant and rapid endochondral bone regeneration using framework-enhanced 3D biomineralized matrix hydrogels. First, it is meticulously designed 3D biomimetic hydrogels with both hypoxic and osteoinductive microenvironment, and then integrated 3D-printed polycaprolactone framework to improve their mechanical strength and structural fidelity. The inherent hypoxic 3D matrix microenvironment effectively activates bone marrow mesenchymal stem cells self-regulation for early-stage chondrogenesis via TGFβ/Smad signaling pathway due to the obstacle of aerobic respiration. Meanwhile, the strong biomineralized microenvironment, created by a hybrid formulation of native-constitute osteogenic inorganic salts, can synergistically regulate both bone mineralization and osteoclastic differentiation, and thus accelerate the late-stage bone maturation. Furthermore, both in vivo ectopic osteogenesis and in situ skull defect repair successfully verified the high efficiency and mechanical maintenance of endochondral bone regeneration mode, which offers a promising treatment for craniofacial bone defect repair.
Keywords: biomineralization; bone regeneration; endochondral ossification; hydrogels; hypoxic microenvironment.
© 2023 The Authors. Advanced Science published by Wiley-VCH GmbH.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Fractionated human adipose tissue as a native biomaterial for the generation of a bone organ by endochondral ossification.Acta Biomater. 2018 Sep 1;77:142-154. doi: 10.1016/j.actbio.2018.07.004. Epub 2018 Jul 4. Acta Biomater. 2018. PMID: 30126590
-
Suppressing mesenchymal stem cell hypertrophy and endochondral ossification in 3D cartilage regeneration with nanofibrous poly(l-lactic acid) scaffold and matrilin-3.Acta Biomater. 2018 Aug;76:29-38. doi: 10.1016/j.actbio.2018.06.027. Epub 2018 Jun 22. Acta Biomater. 2018. PMID: 29940371 Free PMC article.
-
Evaluation of an Engineered Hybrid Matrix for Bone Regeneration via Endochondral Ossification.Ann Biomed Eng. 2020 Mar;48(3):992-1005. doi: 10.1007/s10439-019-02279-0. Epub 2019 Apr 29. Ann Biomed Eng. 2020. PMID: 31037444 Free PMC article.
-
Endochondral ossification for enhancing bone regeneration: converging native extracellular matrix biomaterials and developmental engineering in vivo.Tissue Eng Part B Rev. 2015 Jun;21(3):247-66. doi: 10.1089/ten.TEB.2014.0419. Epub 2014 Dec 4. Tissue Eng Part B Rev. 2015. PMID: 25336144 Free PMC article. Review.
-
Biomimetic Hydrogels as the Inductive Endochondral Ossification Template for Promoting Bone Regeneration.Adv Healthc Mater. 2024 Jul;13(18):e2303532. doi: 10.1002/adhm.202303532. Epub 2024 May 25. Adv Healthc Mater. 2024. PMID: 38108565 Review.
Cited by
-
Screening of molecular markers associated with hornless traits in Qira black sheep.BMC Genomics. 2025 May 8;26(1):458. doi: 10.1186/s12864-025-11608-8. BMC Genomics. 2025. PMID: 40340543 Free PMC article.
-
Fabricating oxygen self-supplying 3D printed bioactive hydrogel scaffold for augmented vascularized bone regeneration.Bioact Mater. 2024 Jun 14;40:227-243. doi: 10.1016/j.bioactmat.2024.06.016. eCollection 2024 Oct. Bioact Mater. 2024. PMID: 38973993 Free PMC article.
-
Rapid neuralized and vascularized osteogenesis in infected bone defect using biomimetic biomineralized and antibacterial hydrogels.Front Bioeng Biotechnol. 2025 May 21;13:1611639. doi: 10.3389/fbioe.2025.1611639. eCollection 2025. Front Bioeng Biotechnol. 2025. PMID: 40470505 Free PMC article.
References
-
- Okuchi Y., Reeves J., Ng S. S., Doro D. H., Junyent S., Liu K. J., El Haj A. J., Habib S. J., Nat. Mater. 2021, 20, 108. - PubMed
-
- An B., Wang Y., Huang Y., Wang X., Liu Y., Xun D., Church G. M., Dai Z., Yi X., Tang T.‐C., Zhong C., Chem. Rev. 2023, 123, 2349. - PubMed
-
- Mao C., Xiao P., Tao X.‐N., Qin J., He Q.‐T., Zhang C., Guo S.‐C., Du Y.‐Q., Chen L.‐N., Shen D.‐D., Yang Z.‐S., Zhang H.‐Q., Huang S.‐M., He Y.‐H., Cheng J., Zhong Y.‐N., Shang P., Chen J., Zhang D.‐L., Wang Q.‐L., Liu M.‐X., Li G.‐Y., Guo Y., Xu H. E., Wang C., Zhang C., Feng S., Yu X., Zhang Y., Sun J.‐P., Science 2023, 380, eadd6220. - PubMed
-
- Zhang Y., Chen M., Dai Z., Cao H., Li J., Zhang W., Biomater. Sci. 2020, 8, 682. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- 2022YFA1207500/National Key Research and Development Program of China
- 81871502/National Natural Science Foundation of China
- 81671837/National Natural Science Foundation of China
- 2022LHA07/Biomaterials and Regenerative Medicine Institute Cooperative Research Project of Shanghai Jiaotong University School of Medicine
- 221111310100/Key Research and Development Program of Henan province
LinkOut - more resources
Full Text Sources
