Control of CCR5 Cell-Surface Targeting by the PRAF2 Gatekeeper
- PMID: 38139265
- PMCID: PMC10744302
- DOI: 10.3390/ijms242417438
Control of CCR5 Cell-Surface Targeting by the PRAF2 Gatekeeper
Abstract
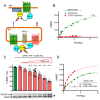
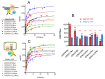
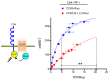
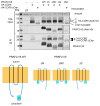
The cell-surface targeting of neo-synthesized G protein-coupled receptors (GPCRs) involves the recruitment of receptors into COPII vesicles budding at endoplasmic reticulum exit sites (ERESs). This process is regulated for some GPCRs by escort proteins, which facilitate their export, or by gatekeepers that retain the receptors in the ER. PRAF2, an ER-resident four trans- membrane domain protein with cytoplasmic extremities, operates as a gatekeeper for the GB1 protomer of the heterodimeric GABAB receptor, interacting with a tandem di-leucine/RXR retention motif in the carboxyterminal tail of GB1. PRAF2 was also reported to interact in a two-hybrid screen with a peptide corresponding to the carboxyterminal tail of the chemokine receptor CCR5 despite the absence of RXR motifs in its sequence. Using a bioluminescence resonance energy transfer (BRET)-based subcellular localization system, we found that PRAF2 inhibits, in a concentration-dependent manner, the plasma membrane export of CCR5. BRET-based proximity assays and Co-IP experiments demonstrated that PRAF2/CCR5 interaction does not require the presence of a receptor carboxyterminal tail and involves instead the transmembrane domains of both proteins. The mutation of the potential di-leucine/RXR motif contained in the third intracellular loop of CCR5 does not affect PRAF2-mediated retention. It instead impairs the cell-surface export of CCR5 by inhibiting CCR5's interaction with its private escort protein, CD4. PRAF2 and CD4 thus display opposite roles on the cell-surface export of CCR5, with PRAF2 inhibiting and CD4 promoting this process, likely operating at the level of CCR5 recruitment into COPII vesicles, which leave the ER.
Keywords: BRET; ERES; G protein-coupled receptor; biosensor; cell trafficking; endoplasmic reticulum; escort protein; gatekeeper.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Pharmacological chaperone-rescued cystic fibrosis CFTR-F508del mutant overcomes PRAF2-gated access to endoplasmic reticulum exit sites.Cell Mol Life Sci. 2022 Sep 27;79(10):530. doi: 10.1007/s00018-022-04554-1. Cell Mol Life Sci. 2022. PMID: 36167862 Free PMC article.
-
GABAB receptor cell-surface export is controlled by an endoplasmic reticulum gatekeeper.Mol Psychiatry. 2016 Apr;21(4):480-90. doi: 10.1038/mp.2015.72. Epub 2015 Jun 2. Mol Psychiatry. 2016. PMID: 26033241 Free PMC article.
-
Gatekeepers Controlling GPCR Export and Function.Trends Pharmacol Sci. 2015 Oct;36(10):636-644. doi: 10.1016/j.tips.2015.06.007. Trends Pharmacol Sci. 2015. PMID: 26435209 Review.
-
Genomic organization, expression profile, and characterization of the new protein PRA1 domain family, member 2 (PRAF2).Gene. 2006 Apr 12;371(1):154-65. doi: 10.1016/j.gene.2005.12.009. Epub 2006 Feb 14. Gene. 2006. PMID: 16481131
-
Regulated GPCR trafficking to the plasma membrane: general issues and the CCR5 chemokine receptor example.Subcell Biochem. 2012;63:97-111. doi: 10.1007/978-94-007-4765-4_6. Subcell Biochem. 2012. PMID: 23161135 Review.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

