PBRM-1/PBAF-regulated genes in a multipotent progenitor in Caenorhabditis elegans
- PMID: 38150396
- PMCID: PMC10917506
- DOI: 10.1093/g3journal/jkad297
PBRM-1/PBAF-regulated genes in a multipotent progenitor in Caenorhabditis elegans
Abstract
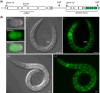
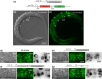
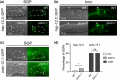
The Caenorhabditis elegans somatic gonadal precursors (SGPs) are multipotent progenitors that generate all somatic cells of the adult reproductive system. The 2 SGPs originate in the mesodermal layer and are born through a division that produces one SGP and one head mesodermal cell (hmc). One hmc terminally differentiates, and the other dies by programmed cell death. The polybromo-associated BAF (PBAF) chromatin remodeling complex promotes the multipotent SGP fate. The complete loss of PBAF causes lethality, so we used a combination of Cre/lox recombination and GFP nanobody-directed protein degradation to eliminate PBRM-1, the signature subunit of the PBAF complex, from 83 mesodermal cells, including SGPs, body muscles, and the hmc. We used RNA sequencing to identify genes acting downstream of PBAF in these cells and identified 1,955 transcripts that were significantly differentially expressed between pbrm-1(-) and pbrm-1(+) in the mesoderm of L1 larvae. We found that genes involved in muscle cell function were overrepresented; most of these genes had lower expression in the absence of PBRM-1, suggesting that PBAF promotes muscle differentiation. Among the differentially expressed genes were 125 that are normally expressed at higher levels in SGP vs hmc and positively regulated by pbrm-1 and 53 that are normally expressed at higher levels in hmc vs SGP and are negatively regulated by pbrm-1; these are candidate regulators of the SGP/hmc fate decision. We validated one candidate gene using a fluorescent reporter; the hsp-12.3 reporter was derepressed in SGPs in pbrm-1 mutants, suggesting that hsp-12.3 expression is normally repressed by pbrm-1 in SGPs.
Keywords: C. elegans; pbrm-1; PBAF; SGP; SWI/SNF chromatin remodeling; differential gene expression; hmc; multipotent progenitor; muscle.
© The Author(s) 2023. Published by Oxford University Press on behalf of The Genetics Society of America.
Conflict of interest statement
Conflicts of interest The author(s) declare no conflicts of interest.
Figures

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous
