Region-Specific Phosphorylation Determines Neuroligin-3 Localization to Excitatory Versus Inhibitory Synapses
- PMID: 38154503
- PMCID: PMC11209832
- DOI: 10.1016/j.biopsych.2023.12.020
Region-Specific Phosphorylation Determines Neuroligin-3 Localization to Excitatory Versus Inhibitory Synapses
Abstract
Background: Neuroligin-3 is a postsynaptic adhesion molecule involved in synapse development and function. It is implicated in rare, monogenic forms of autism, and its shedding is critical to the tumor microenvironment of gliomas. While other members of the neuroligin family exhibit synapse-type specificity in localization and function through distinct interactions with postsynaptic scaffold proteins, the specificity of neuroligin-3 synaptic localization remains largely unknown.
Methods: We investigated the synaptic localization of neuroligin-3 across regions in mouse and human brain samples after validating antibody specificity in knockout animals. We raised a phospho-specific neuroligin antibody and used phosphoproteomics, cell-based assays, and in utero CRISPR/Cas9 (clustered regularly interspaced short palindromic repeats/Cas9) knockout and gene replacement to identify mechanisms that regulate neuroligin-3 localization to distinct synapse types.
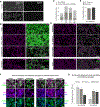
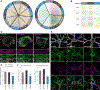
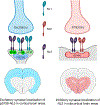
Results: Neuroligin-3 exhibits region-dependent synapse specificity, largely localizing to excitatory synapses in cortical regions and inhibitory synapses in subcortical regions of the brain in both mice and humans. We identified specific phosphorylation of cortical neuroligin-3 at a key binding site for recruitment to inhibitory synapses, while subcortical neuroligin-3 remained unphosphorylated. In vitro, phosphomimetic mutation of that site disrupted neuroligin-3 association with the inhibitory postsynaptic scaffolding protein gephyrin. In vivo, phosphomimetic mutants of neuroligin-3 localized to excitatory postsynapses, while phospho-null mutants localized to inhibitory postsynapses.
Conclusions: These data reveal an unexpected region-specific pattern of neuroligin-3 synapse specificity, as well as a phosphorylation-dependent mechanism that regulates its recruitment to either excitatory or inhibitory synapses. These findings add to our understanding of how neuroligin-3 is involved in conditions that may affect the balance of excitation and inhibition.
Keywords: Autism; Excitatory synapse; Inhibitory synapse; Neuroligin; Phosphorylation; Scaffolding protein.
Copyright © 2023 Society of Biological Psychiatry. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
The authors report no biomedical financial interests or potential conflicts of interest.
Figures
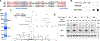


References
-
- Bourgeron T (2015): From the genetic architecture to synaptic plasticity in autism spectrum disorder. Nat. Rev. Neurosci 16 (9): 551–63. - PubMed
-
- Krueger DD, Tuffy LP, Papadopoulos T, Brose N (2012): The role of neurexins and neuroligins in the formation, maturation, and function of vertebrate synapses. Curr. Opin. Neurobiol 22 (3): 412–22. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

