Spatiotemporal modulation of growth factors directs the generation of multilineage mouse embryonic stem cell-derived mammary organoids
- PMID: 38159568
- PMCID: PMC10872289
- DOI: 10.1016/j.devcel.2023.12.003
Spatiotemporal modulation of growth factors directs the generation of multilineage mouse embryonic stem cell-derived mammary organoids
Abstract
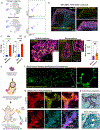
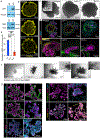
Ectodermal appendages, such as the mammary gland (MG), are thought to have evolved from hair-associated apocrine glands to serve the function of milk secretion. Through the directed differentiation of mouse embryonic stem cells (mESCs), here, we report the generation of multilineage ESC-derived mammary organoids (MEMOs). We adapted the skin organoid model, inducing the dermal mesenchyme to transform into mammary-specific mesenchyme via the sequential activation of Bone Morphogenetic Protein 4 (BMP4) and Parathyroid Hormone-related Protein (PTHrP) and inhibition of hedgehog (HH) signaling. Using single-cell RNA sequencing, we identified gene expression profiles that demonstrate the presence of mammary-specific epithelial cells, fibroblasts, and adipocytes. MEMOs undergo ductal morphogenesis in Matrigel and can reconstitute the MG in vivo. Further, we demonstrate that the loss of function in placode regulators LEF1 and TBX3 in mESCs results in impaired skin and MEMO generation. In summary, our MEMO model is a robust tool for studying the development of ectodermal appendages, and it provides a foundation for regenerative medicine and disease modeling.
Keywords: ectodermal appendages; embryonic stem cell; mammary gland; organoid; single-cell transcriptomics.
Published by Elsevier Inc.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures


Similar articles
-
Embryonic Mammary Gland Morphogenesis.Adv Exp Med Biol. 2025;1464:9-27. doi: 10.1007/978-3-031-70875-6_2. Adv Exp Med Biol. 2025. PMID: 39821018 Review.
-
Lef1 is required for the transition of Wnt signaling from mesenchymal to epithelial cells in the mouse embryonic mammary gland.Dev Biol. 2006 Jul 1;295(1):219-31. doi: 10.1016/j.ydbio.2006.03.030. Epub 2006 Apr 3. Dev Biol. 2006. PMID: 16678815
-
BMP4 and PTHrP interact to stimulate ductal outgrowth during embryonic mammary development and to inhibit hair follicle induction.Development. 2007 Mar;134(6):1221-30. doi: 10.1242/dev.000182. Epub 2007 Feb 14. Development. 2007. PMID: 17301089
-
Parathyroid hormone-related protein maintains mammary epithelial fate and triggers nipple skin differentiation during embryonic breast development.Development. 2001 Feb;128(4):513-25. doi: 10.1242/dev.128.4.513. Development. 2001. PMID: 11171335
-
Parathyroid hormone-related protein specifies the mammary mesenchyme and regulates embryonic mammary development.J Mammary Gland Biol Neoplasia. 2013 Jun;18(2):171-7. doi: 10.1007/s10911-013-9283-7. Epub 2013 May 3. J Mammary Gland Biol Neoplasia. 2013. PMID: 23640717 Free PMC article. Review.
Cited by
-
Spatially defined microenvironment for engineering organoids.Biophys Rev (Melville). 2024 Oct 18;5(4):041302. doi: 10.1063/5.0198848. eCollection 2024 Dec. Biophys Rev (Melville). 2024. PMID: 39679203 Review.
-
From morphology to single-cell molecules: high-resolution 3D histology in biomedicine.Mol Cancer. 2025 Mar 3;24(1):63. doi: 10.1186/s12943-025-02240-x. Mol Cancer. 2025. PMID: 40033282 Free PMC article. Review.
-
Small molecules direct the generation of ameloblast-like cells from human embryonic stem cells.Stem Cell Res Ther. 2025 Apr 12;16(1):173. doi: 10.1186/s13287-025-04294-6. Stem Cell Res Ther. 2025. PMID: 40221796 Free PMC article.
-
Organoid-Guided Precision Medicine: From Bench to Bedside.MedComm (2020). 2025 May 1;6(5):e70195. doi: 10.1002/mco2.70195. eCollection 2025 May. MedComm (2020). 2025. PMID: 40321594 Free PMC article. Review.
-
Embryonic Mammary Gland Morphogenesis.Adv Exp Med Biol. 2025;1464:9-27. doi: 10.1007/978-3-031-70875-6_2. Adv Exp Med Biol. 2025. PMID: 39821018 Review.
References
-
- Oftedal OT (2002). evolutionary origin of mammary gland. Journal of Mammary Gland Biology and Neoplasia 7, 225–252. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

