Segmentation of cellular ultrastructures on sparsely labeled 3D electron microscopy images using deep learning
- PMID: 38162124
- PMCID: PMC10754953
- DOI: 10.3389/fbinf.2023.1308708
Segmentation of cellular ultrastructures on sparsely labeled 3D electron microscopy images using deep learning
Abstract
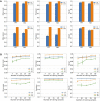
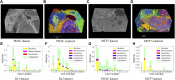
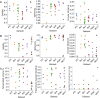
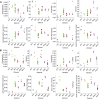
Focused ion beam-scanning electron microscopy (FIB-SEM) images can provide a detailed view of the cellular ultrastructure of tumor cells. A deeper understanding of their organization and interactions can shed light on cancer mechanisms and progression. However, the bottleneck in the analysis is the delineation of the cellular structures to enable quantitative measurements and analysis. We mitigated this limitation using deep learning to segment cells and subcellular ultrastructure in 3D FIB-SEM images of tumor biopsies obtained from patients with metastatic breast and pancreatic cancers. The ultrastructures, such as nuclei, nucleoli, mitochondria, endosomes, and lysosomes, are relatively better defined than their surroundings and can be segmented with high accuracy using a neural network trained with sparse manual labels. Cell segmentation, on the other hand, is much more challenging due to the lack of clear boundaries separating cells in the tissue. We adopted a multi-pronged approach combining detection, boundary propagation, and tracking for cell segmentation. Specifically, a neural network was employed to detect the intracellular space; optical flow was used to propagate cell boundaries across the z-stack from the nearest ground truth image in order to facilitate the separation of individual cells; finally, the filopodium-like protrusions were tracked to the main cells by calculating the intersection over union measure for all regions detected in consecutive images along z-stack and connecting regions with maximum overlap. The proposed cell segmentation methodology resulted in an average Dice score of 0.93. For nuclei, nucleoli, and mitochondria, the segmentation achieved Dice scores of 0.99, 0.98, and 0.86, respectively. The segmentation of FIB-SEM images will enable interpretative rendering and provide quantitative image features to be associated with relevant clinical variables.
Keywords: cell boundary; deep neural network; electron microscopy; segmentation; subcellular ultrastructure.
Copyright © 2023 Machireddy, Thibault, Loftis, Stoltz, Bueno, Smith, Riesterer, Gray and Song.
Conflict of interest statement
JG has licensed technologies to Abbott Diagnostics; has ownership positions in Convergent Genomics, Health Technology Innovations, Zorro Bio, and PDX Pharmaceuticals; serves as a paid consultant to New Leaf Ventures; has received research support from Thermo Fisher Scientific (formerly FEI), Zeiss, Miltenyi Biotec, Quantitative Imaging, Health Technology Innovations, and Micron Technologies; and owns stock in Abbott Diagnostics, AbbVie, Alphabet, Amazon, Amgen, Apple, General Electric, Gilead, Intel, Microsoft, NVIDIA, and Zimmer Biomet. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. The author(s) declared that they were an editorial board member of Frontiers, at the time of submission. This had no impact on the peer review process and the final decision.
Figures








Similar articles
-
An Evaluation of Computational Learning-based Methods for the Segmentation of Nuclei in Cervical Cancer Cells from Microscopic Images.Curr Comput Aided Drug Des. 2022;18(2):81-94. doi: 10.2174/1573409918666220208120756. Curr Comput Aided Drug Des. 2022. PMID: 35139795
-
Automated cell structure extraction for 3D electron microscopy by deep learning.Sci Rep. 2025 May 20;15(1):17481. doi: 10.1038/s41598-025-01763-z. Sci Rep. 2025. PMID: 40394179 Free PMC article.
-
Automated segmentation of cardiomyocyte Z-disks from high-throughput scanning electron microscopy data.BMC Med Inform Decis Mak. 2019 Dec 19;19(Suppl 6):272. doi: 10.1186/s12911-019-0962-1. BMC Med Inform Decis Mak. 2019. PMID: 31856827 Free PMC article.
-
Automatic Segmentation of Multiple Organs on 3D CT Images by Using Deep Learning Approaches.Adv Exp Med Biol. 2020;1213:135-147. doi: 10.1007/978-3-030-33128-3_9. Adv Exp Med Biol. 2020. PMID: 32030668 Review.
-
Deep learning for 3D imaging and image analysis in biomineralization research.J Struct Biol. 2020 Oct 1;212(1):107598. doi: 10.1016/j.jsb.2020.107598. Epub 2020 Aug 9. J Struct Biol. 2020. PMID: 32783967 Review.
Cited by
-
Efficient semi-supervised semantic segmentation of electron microscopy cancer images with sparse annotations.bioRxiv [Preprint]. 2023 Nov 1:2023.10.30.563998. doi: 10.1101/2023.10.30.563998. bioRxiv. 2023. Update in: Front Bioinform. 2023 Dec 15;3:1308707. doi: 10.3389/fbinf.2023.1308707. PMID: 37961180 Free PMC article. Updated. Preprint.
-
How innovations in methodology offer new prospects for volume electron microscopy.J Microsc. 2022 Sep;287(3):114-137. doi: 10.1111/jmi.13134. Epub 2022 Jul 27. J Microsc. 2022. PMID: 35810393 Free PMC article. Review.
-
Online citizen science with the Zooniverse for analysis of biological volumetric data.Histochem Cell Biol. 2023 Sep;160(3):253-276. doi: 10.1007/s00418-023-02204-6. Epub 2023 Jun 7. Histochem Cell Biol. 2023. PMID: 37284846 Free PMC article.
-
Efficient semi-supervised semantic segmentation of electron microscopy cancer images with sparse annotations.Front Bioinform. 2023 Dec 15;3:1308707. doi: 10.3389/fbinf.2023.1308707. eCollection 2023. Front Bioinform. 2023. PMID: 38162122 Free PMC article.
References
-
- Abadi M., Barham P., Chen J., Chen Z., Davis A., Dean J., et al. (2016). “TensorFlow: a system for {Large-Scale} machine learning,” in 12th USENIX symposium on operating systems design and implementation (OSDI 16) (Savannah, GA, USA: ACM; ), 265–283.
-
- Baba A. I., Câtoi C. (2007). “Tumor cell morphology,” in Comparative oncology (Bucuresti, România: The Publishing House of the Romanian Academy; ). - PubMed
-
- Barnes R., Lehman C., Mulla D. (2014). Priority-flood: an optimal depression-filling and watershed-labeling algorithm for digital elevation models. Comput. Geosciences 62, 117–127. 10.1016/j.cageo.2013.04.024 - DOI
Grants and funding
LinkOut - more resources
Full Text Sources

