Overproduction of mycotoxin biosynthetic enzymes triggers Fusarium toxisome-shaped structure formation via endoplasmic reticulum remodeling
- PMID: 38166144
- PMCID: PMC10786393
- DOI: 10.1371/journal.ppat.1011913
Overproduction of mycotoxin biosynthetic enzymes triggers Fusarium toxisome-shaped structure formation via endoplasmic reticulum remodeling
Abstract
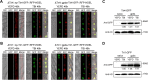
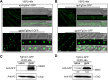
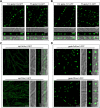
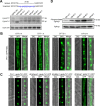
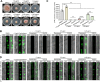
Mycotoxin deoxynivalenol (DON) produced by the Fusarium graminearum complex is highly toxic to animal and human health. During DON synthesis, the endoplasmic reticulum (ER) of F. graminearum is intensively reorganized, from thin reticular structure to thickened spherical and crescent structure, which was referred to as "DON toxisome". However, the underlying mechanism of how the ER is reorganized into toxisome remains unknown. In this study, we discovered that overproduction of ER-localized DON biosynthetic enzyme Tri4 or Tri1, or intrinsic ER-resident membrane proteins FgHmr1 and FgCnx was sufficient to induce toxisome-shaped structure (TSS) formation under non-toxin-inducing conditions. Moreover, heterologous overexpression of Tri1 and Tri4 proteins in non-DON-producing fungi F. oxysporum f. sp. lycopersici and F. fujikuroi also led to TSS formation. In addition, we found that the high osmolarity glycerol (HOG), but not the unfolded protein response (UPR) signaling pathway was involved in the assembly of ER into TSS. By using toxisome as a biomarker, we screened and identified a novel chemical which exhibited high inhibitory activity against toxisome formation and DON biosynthesis, and inhibited Fusarium growth species-specifically. Taken together, this study demonstrated that the essence of ER remodeling into toxisome structure is a response to the overproduction of ER-localized DON biosynthetic enzymes, providing a novel pathway for management of mycotoxin contamination.
Copyright: © 2024 Wang et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

