GCNFORMER: graph convolutional network and transformer for predicting lncRNA-disease associations
- PMID: 38166659
- PMCID: PMC10763317
- DOI: 10.1186/s12859-023-05625-1
GCNFORMER: graph convolutional network and transformer for predicting lncRNA-disease associations
Abstract
Background: A growing body of researches indicate that the disrupted expression of long non-coding RNA (lncRNA) is linked to a range of human disorders. Therefore, the effective prediction of lncRNA-disease association (LDA) can not only suggest solutions to diagnose a condition but also save significant time and labor costs.
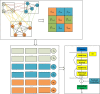
Method: In this work, we proposed a novel LDA predicting algorithm based on graph convolutional network and transformer, named GCNFORMER. Firstly, we integrated the intraclass similarity and interclass connections between miRNAs, lncRNAs and diseases, and built a graph adjacency matrix. Secondly, to completely obtain the features between various nodes, we employed a graph convolutional network for feature extraction. Finally, to obtain the global dependencies between inputs and outputs, we used a transformer encoder with a multiheaded attention mechanism to forecast lncRNA-disease associations.
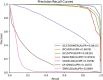
Results: The results of fivefold cross-validation experiment on the public dataset revealed that the AUC and AUPR of GCNFORMER achieved 0.9739 and 0.9812, respectively. We compared GCNFORMER with six advanced LDA prediction models, and the results indicated its superiority over the other six models. Furthermore, GCNFORMER's effectiveness in predicting potential LDAs is underscored by case studies on breast cancer, colon cancer and lung cancer.
Conclusions: The combination of graph convolutional network and transformer can effectively improve the performance of LDA prediction model and promote the in-depth development of this research filed.
Keywords: Graph convolutional network; LncRNA-disease association prediction; Machine learning; Multiheaded attention mechanism; transformer.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures
Similar articles
-
Node-adaptive graph Transformer with structural encoding for accurate and robust lncRNA-disease association prediction.BMC Genomics. 2024 Jan 18;25(1):73. doi: 10.1186/s12864-024-09998-2. BMC Genomics. 2024. PMID: 38233788 Free PMC article.
-
Graph Convolutional Network and Convolutional Neural Network Based Method for Predicting lncRNA-Disease Associations.Cells. 2019 Aug 30;8(9):1012. doi: 10.3390/cells8091012. Cells. 2019. PMID: 31480350 Free PMC article.
-
LDAGM: prediction lncRNA-disease asociations by graph convolutional auto-encoder and multilayer perceptron based on multi-view heterogeneous networks.BMC Bioinformatics. 2024 Oct 15;25(1):332. doi: 10.1186/s12859-024-05950-z. BMC Bioinformatics. 2024. PMID: 39407120 Free PMC article.
-
Recent advances in machine learning methods for predicting LncRNA and disease associations.Front Cell Infect Microbiol. 2022 Nov 30;12:1071972. doi: 10.3389/fcimb.2022.1071972. eCollection 2022. Front Cell Infect Microbiol. 2022. PMID: 36530425 Free PMC article. Review.
-
Data resources and computational methods for lncRNA-disease association prediction.Comput Biol Med. 2023 Feb;153:106527. doi: 10.1016/j.compbiomed.2022.106527. Epub 2023 Jan 2. Comput Biol Med. 2023. PMID: 36610216 Review.
Cited by
-
RNA sequence analysis landscape: A comprehensive review of task types, databases, datasets, word embedding methods, and language models.Heliyon. 2025 Jan 6;11(2):e41488. doi: 10.1016/j.heliyon.2024.e41488. eCollection 2025 Jan 30. Heliyon. 2025. PMID: 39897847 Free PMC article. Review.
-
Neighborhood based computational approaches for the prediction of lncRNA-disease associations.BMC Bioinformatics. 2024 May 13;25(1):187. doi: 10.1186/s12859-024-05777-8. BMC Bioinformatics. 2024. PMID: 38741200 Free PMC article.
-
HGATLink: single-cell gene regulatory network inference via the fusion of heterogeneous graph attention networks and transformer.BMC Bioinformatics. 2025 Feb 11;26(1):49. doi: 10.1186/s12859-025-06071-x. BMC Bioinformatics. 2025. PMID: 39934680 Free PMC article.
-
GL4SDA: Predicting snoRNA-disease associations using GNNs and LLM embeddings.Comput Struct Biotechnol J. 2025 Mar 12;27:1023-1033. doi: 10.1016/j.csbj.2025.03.014. eCollection 2025. Comput Struct Biotechnol J. 2025. PMID: 40160859 Free PMC article.
References
-
- Djebali S, Davis CA, Merkel A, Dobin A, Lassmann T, Mortazavi A, Tanzer A, Lagarde J, Lin W, Schlesinger F, Xue C, Marinov GK, Khatun J, Williams BA, Zaleski C, Rozowsky J, Röder M, Kokocinski F, Abdelhamid RF, Alioto T, Gingeras TR. Landscape of transcription in human cells. Nature. 2012;489(7414):101–108. doi: 10.1038/nature11233. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical