The evolution of centriole degradation in mouse sperm
- PMID: 38168044
- PMCID: PMC10761967
- DOI: 10.1038/s41467-023-44411-8
The evolution of centriole degradation in mouse sperm
Abstract
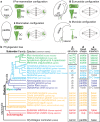
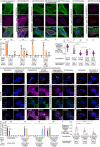
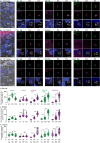
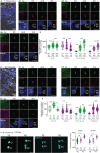
Centrioles are subcellular organelles found at the cilia base with an evolutionarily conserved structure and a shock absorber-like function. In sperm, centrioles are found at the flagellum base and are essential for embryo development in basal animals. Yet, sperm centrioles have evolved diverse forms, sometimes acting like a transmission system, as in cattle, and sometimes becoming dispensable, as in house mice. How the essential sperm centriole evolved to become dispensable in some organisms is unclear. Here, we test the hypothesis that this transition occurred through a cascade of evolutionary changes to the proteins, structure, and function of sperm centrioles and was possibly driven by sperm competition. We found that the final steps in this cascade are associated with a change in the primary structure of the centriolar inner scaffold protein FAM161A in rodents. This information provides the first insight into the molecular mechanisms and adaptive evolution underlying a major evolutionary transition within the internal structure of the mammalian sperm neck.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



References
-
- Parker GA. Sperm competition and its evolutionary consequences in the insects. Biol. Rev. 1970;45:525–567. doi: 10.1111/j.1469-185X.1970.tb01176.x. - DOI
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

