This is a preprint.
Hybracter: Enabling Scalable, Automated, Complete and Accurate Bacterial Genome Assemblies
- PMID: 38168369
- PMCID: PMC10760025
- DOI: 10.1101/2023.12.12.571215
Hybracter: Enabling Scalable, Automated, Complete and Accurate Bacterial Genome Assemblies
Update in
-
Hybracter: enabling scalable, automated, complete and accurate bacterial genome assemblies.Microb Genom. 2024 May;10(5):001244. doi: 10.1099/mgen.0.001244. Microb Genom. 2024. PMID: 38717808 Free PMC article.
Abstract
Improvements in the accuracy and availability of long-read sequencing mean that complete bacterial genomes are now routinely reconstructed using hybrid (i.e. short- and long-reads) assembly approaches. Complete genomes allow a deeper understanding of bacterial evolution and genomic variation beyond single nucleotide variants (SNVs). They are also crucial for identifying plasmids, which often carry medically significant antimicrobial resistance (AMR) genes. However, small plasmids are often missed or misassembled by long-read assembly algorithms. Here, we present Hybracter which allows for the fast, automatic, and scalable recovery of near-perfect complete bacterial genomes using a long-read first assembly approach. Hybracter can be run either as a hybrid assembler or as a long-read only assembler. We compared Hybracter to existing automated hybrid and long-read only assembly tools using a diverse panel of samples of varying levels of long-read accuracy with manually curated ground truth reference genomes. We demonstrate that Hybracter as a hybrid assembler is more accurate and faster than the existing gold standard automated hybrid assembler Unicycler. We also show that Hybracter with long-reads only is the most accurate long-read only assembler and is comparable to hybrid methods in accurately recovering small plasmids.
Conflict of interest statement
Conflicts of Interest The authors declare that there are no conflicts of interest.
Figures


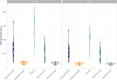

References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
