VAP spatially stabilizes dendritic mitochondria to locally support synaptic plasticity
- PMID: 38177103
- PMCID: PMC10766606
- DOI: 10.1038/s41467-023-44233-8
VAP spatially stabilizes dendritic mitochondria to locally support synaptic plasticity
Abstract
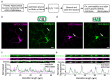
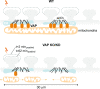
Synapses are pivotal sites of plasticity and memory formation. Consequently, synapses are energy consumption hotspots susceptible to dysfunction when their energy supplies are perturbed. Mitochondria are stabilized near synapses via the cytoskeleton and provide the local energy required for synaptic plasticity. However, the mechanisms that tether and stabilize mitochondria to support synaptic plasticity are unknown. We identified proteins exclusively tethering mitochondria to actin near postsynaptic spines. We find that VAP, the vesicle-associated membrane protein-associated protein implicated in amyotrophic lateral sclerosis, stabilizes mitochondria via actin near the spines. To test if the VAP-dependent stable mitochondrial compartments can locally support synaptic plasticity, we used two-photon glutamate uncaging for spine plasticity induction and investigated the induced and adjacent uninduced spines. We find VAP functions as a spatial stabilizer of mitochondrial compartments for up to ~60 min and as a spatial ruler determining the ~30 μm dendritic segment supported during synaptic plasticity.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures





Similar articles
-
Spatially Stable Mitochondrial Compartments Fuel Local Translation during Plasticity.Cell. 2019 Jan 10;176(1-2):73-84.e15. doi: 10.1016/j.cell.2018.12.013. Epub 2019 Jan 3. Cell. 2019. PMID: 30612742
-
Phosphoinositide-dependent enrichment of actin monomers in dendritic spines regulates synapse development and plasticity.J Cell Biol. 2017 Aug 7;216(8):2551-2564. doi: 10.1083/jcb.201612042. Epub 2017 Jun 28. J Cell Biol. 2017. PMID: 28659327 Free PMC article.
-
Role of Drebrin in Synaptic Plasticity.Adv Exp Med Biol. 2017;1006:183-201. doi: 10.1007/978-4-431-56550-5_11. Adv Exp Med Biol. 2017. PMID: 28865021 Review.
-
Actin in dendritic spines: connecting dynamics to function.J Cell Biol. 2010 May 17;189(4):619-29. doi: 10.1083/jcb.201003008. Epub 2010 May 10. J Cell Biol. 2010. PMID: 20457765 Free PMC article. Review.
-
New waves in dendritic spine actin cytoskeleton: From branches and bundles to rings, from actin binding proteins to post-translational modifications.Mol Cell Neurosci. 2017 Oct;84:77-84. doi: 10.1016/j.mcn.2017.05.002. Epub 2017 May 4. Mol Cell Neurosci. 2017. PMID: 28479292 Review.
Cited by
-
Activation of the 5-HT1A Receptor by Eltoprazine Restores Mitochondrial and Motor Deficits in a Drosophila Model of Fragile X Syndrome.Int J Mol Sci. 2024 Aug 13;25(16):8787. doi: 10.3390/ijms25168787. Int J Mol Sci. 2024. PMID: 39201473 Free PMC article.
-
Reduced DJ-1-F1Fo ATP synthase association correlates with midbrain dopaminergic neuron vulnerability in idiopathic Parkinson's disease.Sci Adv. 2025 Jun 6;11(23):eads3051. doi: 10.1126/sciadv.ads3051. Epub 2025 Jun 6. Sci Adv. 2025. PMID: 40479058 Free PMC article.
-
Structural and functional studies of the VAPB-PTPIP51 ER-mitochondria tethering proteins in neurodegenerative diseases.Acta Neuropathol Commun. 2025 Mar 5;13(1):49. doi: 10.1186/s40478-025-01964-7. Acta Neuropathol Commun. 2025. PMID: 40045432 Free PMC article. Review.
-
Calcium signaling in postsynaptic mitochondria: mechanisms, dynamics, and role in ATP production.Front Mol Neurosci. 2025 Jul 21;18:1621070. doi: 10.3389/fnmol.2025.1621070. eCollection 2025. Front Mol Neurosci. 2025. PMID: 40761376 Free PMC article. Review.
-
ALS-Linked VapB P56S Mutation Alters Neuronal Mitochondrial Turnover at the Synapse.J Neurosci. 2024 Aug 28;44(35):e0879242024. doi: 10.1523/JNEUROSCI.0879-24.2024. J Neurosci. 2024. PMID: 39054069 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

