Phylogenomics analysis of velvet regulators in the fungal kingdom
- PMID: 38179919
- PMCID: PMC10845976
- DOI: 10.1128/spectrum.03717-23
Phylogenomics analysis of velvet regulators in the fungal kingdom
Abstract
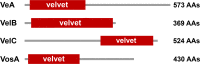
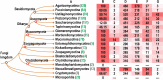
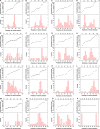
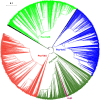
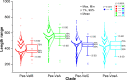
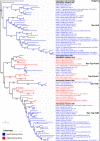
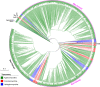
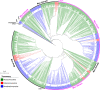
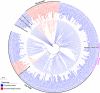
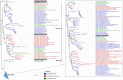
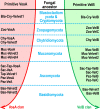
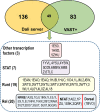
All life forms have evolved to respond appropriately to various environmental and internal cues. In the animal kingdom, the prototypical regulator class of such cellular responses is the Rel homology domain proteins including nuclear factor kappa-light-chain-enhancer of activated B cells (NF-κB). Fungi, the close relatives of animals, have also evolved with their own NF-κB-like regulators called velvet family proteins to govern cellular and chemical development. Here, we conducted a detailed investigation of the taxonomic broad presence of velvet proteins. We observed that velvet proteins are widely distributed in the fungal kingdom. Moreover, we have identified and characterized 21 major velvet clades in fungi. We have further revealed that the highly conserved velvet domain is composed of three distinct motifs and acts as an evolutionarily independent domain, which can be shuffled with various functional domains. Such rearrangements of the velvet domain have resulted in the functional and type diversity of the present velvet regulators. Importantly, our in-deep analyses of the primary and 3D structures of the various velvet domains showed that the fungal velvet domains can be divided into two major clans: the VelB and the VosA clans. The 3D structure comparisons revealed a close similarity of the velvet domain with many other eukaryotic DNA-binding proteins, including those of the Rel, Runt, and signal transducer and activator of transcription families, sharing a common β-sandwich fold. Altogether, this study improves our understanding of velvet regulators in the fungal kingdom.IMPORTANCEFungi are the relatives of animals in Opisthokonta and closely associated with human life by interactive ways such as pathogenicity, food, and secondary metabolites including beneficial ones like penicillin and harmful ones like the carcinogenic aflatoxins. Similar to animals, fungi have also evolved with NF-κB-like velvet family regulators. The velvet proteins constitute a large protein family of fungal transcription factors sharing a common velvet domain and play a key role in coordinating fungal secondary metabolism, developmental and differentiation processes. Our current understanding on velvet regulators is mostly from Ascomycota fungi; however, they remain largely unknown outside Ascomycota. Therefore, this study performed a taxonomic broad investigation of velvet proteins across the fungal kingdom and conducted a detailed analysis on velvet distribution, structure, diversity, and evolution. The results provide a holistic view of velvet regulatory system in the fungal kingdom.
Keywords: DNA-binding; Rel homology domain; fungal development; fungi; nuclear factor kappa-light-chain-enhancer of activated B cells (NF-κB); secondary metabolism; velvet regulators.
Conflict of interest statement
The authors declare no conflict of interest.
Figures






Similar articles
-
Resolving the fungal velvet domain architecture by Aspergillus nidulans VelB.mBio. 2025 May 14;16(5):e0026125. doi: 10.1128/mbio.00261-25. Epub 2025 Mar 31. mBio. 2025. PMID: 40162796 Free PMC article.
-
The velvet family of fungal regulators contains a DNA-binding domain structurally similar to NF-κB.PLoS Biol. 2013 Dec;11(12):e1001750. doi: 10.1371/journal.pbio.1001750. Epub 2013 Dec 31. PLoS Biol. 2013. PMID: 24391470 Free PMC article.
-
Characterization of the velvet regulators in Aspergillus flavus.J Microbiol. 2018 Dec;56(12):893-901. doi: 10.1007/s12275-018-8417-4. Epub 2018 Oct 25. J Microbiol. 2018. PMID: 30361976
-
[Progress in the study of Velvet and LaeA proteins and their relation to the development and bioactive compounds in medicinal fungi].Yao Xue Xue Bao. 2014 Nov;49(11):1520-7. Yao Xue Xue Bao. 2014. PMID: 25757276 Review. Chinese.
-
Coordination of secondary metabolism and development in fungi: the velvet family of regulatory proteins.FEMS Microbiol Rev. 2012 Jan;36(1):1-24. doi: 10.1111/j.1574-6976.2011.00285.x. Epub 2011 Jul 13. FEMS Microbiol Rev. 2012. PMID: 21658084 Review.
Cited by
-
Resolving the fungal velvet domain architecture by Aspergillus nidulans VelB.mBio. 2025 May 14;16(5):e0026125. doi: 10.1128/mbio.00261-25. Epub 2025 Mar 31. mBio. 2025. PMID: 40162796 Free PMC article.
-
Velvet Family Protein FpVelB Affects Virulence in Association with Secondary Metabolism in Fusarium pseudograminearum.Cells. 2024 May 30;13(11):950. doi: 10.3390/cells13110950. Cells. 2024. PMID: 38891082 Free PMC article.
-
Involvement of LaeA and Velvet Proteins in Regulating the Production of Mycotoxins and Other Fungal Secondary Metabolites.J Fungi (Basel). 2024 Aug 8;10(8):561. doi: 10.3390/jof10080561. J Fungi (Basel). 2024. PMID: 39194887 Free PMC article. Review.
-
Genome-wide identification and expression analysis revealed key transcription factors as potential regulators of high-temperature adaptation of Coriolopsis trogii.Arch Microbiol. 2024 Jul 19;206(8):357. doi: 10.1007/s00203-024-04081-6. Arch Microbiol. 2024. PMID: 39028428
References
-
- Wingfield JC, Patrick Kelley J, Angelier F, Chastel O, Lei F, Lynn SE, Miner B, Davis JE, Li D, Wang G. 2011. Organism–environment interactions in a changing world: a mechanistic approach. J Ornithol 152:279–288. doi:10.1007/s10336-011-0668-3 - DOI
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

