Exosomal hsa-miR-151a-3p and hsa-miR-877-5p are potential novel biomarkers for predicting bone metastasis in lung cancer
- PMID: 38180107
- PMCID: PMC10781484
- DOI: 10.18632/aging.205314
Exosomal hsa-miR-151a-3p and hsa-miR-877-5p are potential novel biomarkers for predicting bone metastasis in lung cancer
Abstract
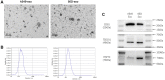
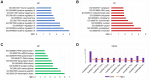
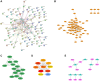
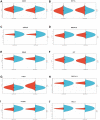
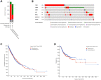
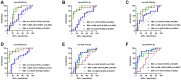
Exosomal miRNAs (exo-miRNAs) have arisen as novel diagnostic biomarkers for various cancers. However, few reports on exo-miRNAs related to bone metastasis (BM) in lung cancer exist. This study aims to screen out key exo-miRNAs and estimate their prognostic values for predicting BM in lung cancer. The differentially expressed exo-miRNAs between the highly-metastatic (95D) and lowly-metastatic (A549) human lung cancer cell lines were comprehensively analyzed using high-throughput sequencing followed by bioinformatic analyses. 29 candidate exo-miRNAs were identified, and 101 BM-related target genes were predicted. Enrichment analysis revealed that these target genes were mainly involved in regulating transcription and pathways in cancer. An exosomal miRNA-mRNA regulatory network consisting of 7 key miRNAs and 10 hub genes was constructed. Further function analysis indicated that these 10 hub genes were mainly enriched in regulating cancer's apoptosis and central carbon metabolism. The survival analysis indicated that 7 of 10 hub genes were closely related to prognosis. Mutation analysis showed that lung cancer patients presented certain genetic alterations in the 7 real hub genes. GSEA for a single hub gene suggested that 6 of 7 real hub genes had close associations with lung cancer development. Finally, ROC analysis revealed that hsa-miR-151a-3p and hsa-miR-877-5p provided high diagnostic accuracy in discriminating patients with bone metastasis (BM+) from patients without bone metastasis (BM-). These findings provided a comprehensive analysis of exo-miRNAs and target genes in the regulatory network of BM in lung cancer. In particular, hsa-miR-151a-3p and hsa-miR-877-5p may be novel biomarkers for predicting BM in lung cancer.
Keywords: biomarker; bone metastasis; exosome; lung cancer; miRNAs.
Conflict of interest statement
Figures





References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

