Intein-based thermoregulated meganucleases for containment of genetic material
- PMID: 38180814
- PMCID: PMC10899782
- DOI: 10.1093/nar/gkad1247
Intein-based thermoregulated meganucleases for containment of genetic material
Abstract
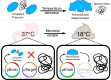
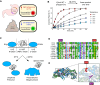
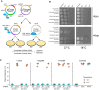
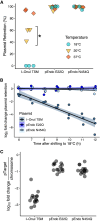
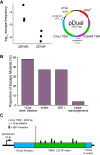
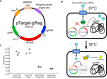
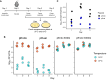
Limiting the spread of synthetic genetic information outside of the intended use is essential for applications where biocontainment is critical. In particular, biocontainment of engineered probiotics and plasmids that are excreted from the mammalian gastrointestinal tract is needed to prevent escape and acquisition of genetic material that could confer a selective advantage to microbial communities. Here, we built a simple and lightweight biocontainment system that post-translationally activates a site-specific DNA endonuclease to degrade DNA at 18°C and not at higher temperatures. We constructed an orthogonal set of temperature-sensitive meganucleases (TSMs) by inserting the yeast VMA1 L212P temperature-sensitive intein into the coding regions of LAGLIDADG homing endonucleases. We showed that the TSMs eliminated plasmids carrying the cognate TSM target site from laboratory strains of Escherichia coli at the permissive 18°C but not at higher restrictive temperatures. Plasmid elimination is dependent on both TSM endonuclease activity and intein splicing. TSMs eliminated plasmids from E. coli Nissle 1917 after passage through the mouse gut when fecal resuspensions were incubated at 18°C but not at 37°C. Collectively, our data demonstrates the potential of thermoregulated meganucleases as a means of restricting engineered plasmids and probiotics to the mammalian gut.
© The Author(s) 2024. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures
Similar articles
-
Structural stability and endonuclease activity of a PI-SceI GFP-fusion protein.Int J Biol Sci. 2007 Feb 16;3(4):205-11. doi: 10.7150/ijbs.3.205. Int J Biol Sci. 2007. PMID: 17389927 Free PMC article.
-
Temperature-sensitive control of protein activity by conditionally splicing inteins.Nat Biotechnol. 2004 Jul;22(7):871-6. doi: 10.1038/nbt979. Epub 2004 Jun 6. Nat Biotechnol. 2004. PMID: 15184905
-
Degeneration of a homing endonuclease and its target sequence in a wild yeast strain.Nucleic Acids Res. 2001 Oct 15;29(20):4215-23. doi: 10.1093/nar/29.20.4215. Nucleic Acids Res. 2001. PMID: 11600710 Free PMC article.
-
Reflections on protein splicing: structures, functions and mechanisms.Proc Jpn Acad Ser B Phys Biol Sci. 2009;85(9):409-21. doi: 10.2183/pjab.85.409. Proc Jpn Acad Ser B Phys Biol Sci. 2009. PMID: 19907126 Free PMC article. Review.
-
The nuclear-encoded inteins of fungi.Fungal Genet Biol. 2007 Mar;44(3):153-79. doi: 10.1016/j.fgb.2006.07.012. Epub 2006 Oct 13. Fungal Genet Biol. 2007. PMID: 17046294 Review.
Cited by
-
A genetic safeguard for eliminating target genes in synthetic probiotics in a gut environment.iScience. 2025 Jun 27;28(8):113027. doi: 10.1016/j.isci.2025.113027. eCollection 2025 Aug 15. iScience. 2025. PMID: 40686611 Free PMC article.
-
Bacterial Species in Engineered Living Materials: Strategies and Future Directions.Microb Biotechnol. 2025 May;18(5):e70164. doi: 10.1111/1751-7915.70164. Microb Biotechnol. 2025. PMID: 40407296 Free PMC article. Review.
References
-
- Lee J.W., Chan C.T., Slomovic S., Collins J.J. Next-generation biocontainment systems for engineered organisms. Nat. Chem. Biol. 2018; 14:530–537. - PubMed
-
- Schmidt M., de Lorenzo V. Synthetic bugs on the loose: containment options for deeply engineered (micro) organisms. Curr. Opin. Biotechnol. 2016; 38:90–96. - PubMed
-
- Isabella V.M., Ha B.N., Castillo M.J., Lubkowicz D.J., Rowe S.E., Millet Y.A., Anderson C.L., Li N., Fisher A.B., West K.A. et al. . Development of a synthetic live bacterial therapeutic for the human metabolic disease phenylketonuria. Nat. Biotechnol. 2018; 36:857–864. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

