SHMT2 Mediates Small-Molecule-Induced Alleviation of Alzheimer Pathology Via the 5'UTR-dependent ADAM10 Translation Initiation
- PMID: 38183387
- PMCID: PMC10953581
- DOI: 10.1002/advs.202305260
SHMT2 Mediates Small-Molecule-Induced Alleviation of Alzheimer Pathology Via the 5'UTR-dependent ADAM10 Translation Initiation
Abstract
It is long been suggested that one-carbon metabolism (OCM) is associated with Alzheimer's disease (AD), whereas the potential mechanisms remain poorly understood. Taking advantage of chemical biology, that mitochondrial serine hydroxymethyltransferase (SHMT2) directly regulated the translation of ADAM metallopeptidase domain 10 (ADAM10), a therapeutic target for AD is reported. That the small-molecule kenpaullone (KEN) promoted ADAM10 translation via the 5' untranslated region (5'UTR) and improved cognitive functions in APP/PS1 mice is found. SHMT2, which is identified as a target gene of KEN and the 5'UTR-interacting RNA binding protein (RBP), mediated KEN-induced ADAM10 translation in vitro and in vivo. SHMT2 controls AD signaling pathways through binding to a large number of RNAs and enhances the 5'UTR activity of ADAM10 by direct interaction with GAGGG motif, whereas this motif affected ribosomal scanning of eukaryotic initiation factor 2 (eIF2) in the 5'UTR. Together, KEN exhibits therapeutic potential for AD by linking OCM with RNA processing, in which the metabolic enzyme SHMT2 "moonlighted" as RBP by binding to GAGGG motif and promoting the 5'UTR-dependent ADAM10 translation initiation.
Keywords: 5′UTR; ADAM10; Alzheimer's disease; Kenpaullone; RNA binding protein; SHMT2; translation inhibitory element.
© 2023 The Authors. Advanced Science published by Wiley-VCH GmbH.
Conflict of interest statement
The authors declare no conflict of interest.
Figures







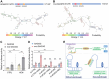
References
-
- a) Nativio R., Lan Y., Donahue G., Sidoli S., Berson A., Srinivasan A. R., Shcherbakova O., Amlie‐Wolf A., Nie J., Cui X., He C., Wang L.‐S., Garcia B. A., Trojanowski J. Q., Bonini N. M., Berger S. L., Nat. Genet. 2020, 52, 1024; - PMC - PubMed
- b) Wang M., Li A., Sekiya M., Beckmann N. D., Quan X., Schrode N., Fernando M. B., Yu A., Zhu L., Cao J., Lyu L., Horgusluoglu E., Wang Q., Guo L., Wang Y.‐S., Neff R., Song W.‐M., Wang E., Shen Q., Zhou X., Ming C., Ho S.‐M., Vatansever S., Kaniskan H. Ü., Jin J., Zhou M.‐M., Ando K., Ho L., Slesinger P. A., Yue Z., et al., Neuron 2021, 109, 257. - PMC - PubMed
-
- Yang J., Long Y., Xu D.‐M., Zhu B.‐L., Deng X.‐J., Yan Z., Sun F., Chen G.‐J., J. Mol. Neurosci. 2019, 69, 608. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
