Non-IG::MYC in diffuse large B-cell lymphoma confers variable genomic configurations and MYC transactivation potential
- PMID: 38184753
- PMCID: PMC10912016
- DOI: 10.1038/s41375-023-02134-1
Non-IG::MYC in diffuse large B-cell lymphoma confers variable genomic configurations and MYC transactivation potential
Abstract
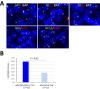
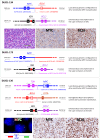
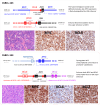
MYC translocation occurs in 8-14% of diffuse large B-cell lymphoma (DLBCL), and may concur with BCL2 and/or BCL6 translocation, known as double-hit (DH) or triple-hit (TH). DLBCL-MYC/BCL2-DH/TH are largely germinal centre B-cell like subtype, but show variable clinical outcome, with IG::MYC fusion significantly associated with inferior survival. While DLBCL-MYC/BCL6-DH are variable in their cell-of-origin subtypes and clinical outcome. Intriguingly, only 40-50% of DLBCL with MYC translocation show high MYC protein expression (>70%). We studied 186 DLBCLs with MYC translocation including 32 MYC/BCL2/BCL6-TH, 75 MYC/BCL2-DH and 26 MYC/BCL6-DH. FISH revealed a MYC/BCL6 fusion in 59% of DLBCL-MYC/BCL2/BCL6-TH and 27% of DLBCL-MYC/BCL6-DH. Targeted NGS showed a similar mutation profile and LymphGen genetic subtype between DLBCL-MYC/BCL2/BCL6-TH and DLBCL-MYC/BCL2-DH, but variable LymphGen subtypes among DLBCL-MYC/BCL6-DH. MYC protein expression is uniformly high in DLBCL with IG::MYC, but variable in those with non-IG::MYC including MYC/BCL6-fusion. Translocation breakpoint analyses of 8 cases by TLC-based NGS showed no obvious genomic configuration that enables MYC transactivation in 3 of the 4 cases with non-IG::MYC, while a typical promoter substitution or IGH super enhancer juxtaposition in the remaining cases. The findings potentially explain variable MYC expression in DLBCL with MYC translocation, and also bear practical implications in its routine assessment.
© 2024. The Author(s).
Conflict of interest statement
ES, JFS and HF are employees of Cergentis, which owns patents on the TLC-NGS method. The authors declare no conflict of interest.
Figures




References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

