This is a preprint.
Spatial analysis of NOS2 and COX2 interaction with T-effector cells reveals immunosuppressive landscapes associated with poor outcome in ER- breast cancer patients
- PMID: 38187660
- PMCID: PMC10769421
- DOI: 10.1101/2023.12.21.572867
Spatial analysis of NOS2 and COX2 interaction with T-effector cells reveals immunosuppressive landscapes associated with poor outcome in ER- breast cancer patients
Abstract
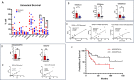
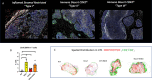
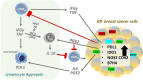
Multiple immunosuppressive mechanisms exist in the tumor microenvironment that drive poor outcomes and decrease treatment efficacy. The co-expression of NOS2 and COX2 is a strong predictor of poor prognosis in ER- breast cancer and other malignancies. Together, they generate pro-oncogenic signals that drive metastasis, drug resistance, cancer stemness, and immune suppression. Using an ER- breast cancer patient cohort, we found that the spatial expression patterns of NOS2 and COX2 with CD3+CD8+PD1- T effector (Teff) cells formed a tumor immune landscape that correlated with poor outcome. NOS2 was primarily associated with the tumor-immune interface, whereas COX2 was associated with immune desert regions of the tumor lacking Teff cells. A higher ratio of NOS2 or COX2 to Teff was highly correlated with poor outcomes. Spatial analysis revealed that regional clustering of NOS2 and COX2 was associated with stromal-restricted Teff, while only COX2 was predominant in immune deserts. Examination of other immunosuppressive elements, such as PDL1/PD1, Treg, B7H4, and IDO1, revealed that PDL1/PD1, Treg, and IDO1 were primarily associated with restricted Teff, whereas B7H4 and COX2 were found in tumor immune deserts. Regardless of the survival outcome, other leukocytes, such as CD4 T cells and macrophages, were primarily in stromal lymphoid aggregates. Finally, in a 4T1 model, COX2 inhibition led to a massive cell infiltration, thus validating the hypothesis that COX2 is an essential component of the Teff exclusion process and, thus, tumor evasion. Our study indicates that NOS2/COX2 expression plays a central role in tumor immunosuppression. Our findings indicate that new strategies combining clinically available NOS2/COX2 inhibitors with various forms of immune therapy may open a new avenue for the treatment of aggressive ER-breast cancers.
Keywords: Breast; COX2; Cancer; NOS2; immunosuppression; spatial.
Figures









Similar articles
-
Elevated tumor NOS2/COX2 promotes immunosuppressive phenotypes associated with poor survival in ER- breast cancer.JCI Insight. 2025 Jul 15;10(16):e193091. doi: 10.1172/jci.insight.193091. eCollection 2025 Aug 22. JCI Insight. 2025. PMID: 40663402
-
NOS2 and COX2 Provide Key Spatial Targets that Determine Outcome in ER- Breast Cancer.bioRxiv [Preprint]. 2023 Dec 23:2023.12.21.572859. doi: 10.1101/2023.12.21.572859. bioRxiv. 2023. PMID: 38187532 Free PMC article. Preprint.
-
Systemic Nos2 Depletion and Cox inhibition limits TNBC disease progression and alters lymphoid cell spatial orientation and density.Redox Biol. 2022 Dec;58:102529. doi: 10.1016/j.redox.2022.102529. Epub 2022 Nov 9. Redox Biol. 2022. PMID: 36375380 Free PMC article.
-
NOS2 and COX-2 Co-Expression Promotes Cancer Progression: A Potential Target for Developing Agents to Prevent or Treat Highly Aggressive Breast Cancer.Int J Mol Sci. 2024 Jun 1;25(11):6103. doi: 10.3390/ijms25116103. Int J Mol Sci. 2024. PMID: 38892290 Free PMC article. Review.
-
The role of the adenosinergic pathway in immunosuppression mediated by human regulatory T cells (Treg).Curr Med Chem. 2011;18(34):5217-23. doi: 10.2174/092986711798184334. Curr Med Chem. 2011. PMID: 22087822 Free PMC article. Review.
Cited by
-
NOS2 as a prognostic biomarker for early-onset colorectal cancer based on public data and clinical validation analysis.Sci Rep. 2025 Feb 4;15(1):4300. doi: 10.1038/s41598-025-88966-6. Sci Rep. 2025. PMID: 39905237 Free PMC article.
References
-
- Vajari M. K., Sanaei M. J., Salari S., Rezvani A., Ravari M. S., and Bashash D. (2023) Breast cancer vaccination: Latest advances with an analytical focus on clinical trials. Int Immunopharmacol 123, 110696. - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
