Glucocorticoid- and pioglitazone-induced proteinuria reduction in experimental NS both correlate with glomerular ECM modulation
- PMID: 38188512
- PMCID: PMC10770536
- DOI: 10.1016/j.isci.2023.108631
Glucocorticoid- and pioglitazone-induced proteinuria reduction in experimental NS both correlate with glomerular ECM modulation
Abstract
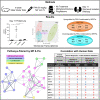
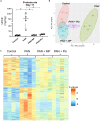
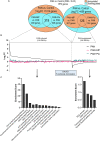
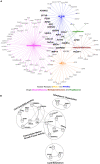
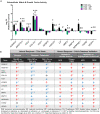
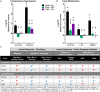
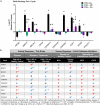
Idiopathic nephrotic syndrome (NS) is a common glomerular disease. Although glucocorticoids (GC) are the primary treatment, the PPARγ agonist pioglitazone (Pio) also reduces proteinuria in patients with NS and directly protects podocytes from injury. Because both drugs reduce proteinuria, we hypothesized these effects result from overlapping transcriptional patterns. Systems biology approaches compared glomerular transcriptomes from rats with PAN-induced NS treated with GC vs. Pio and identified 29 commonly regulated genes-of-interest, primarily involved in extracellular matrix (ECM) remodeling. Correlation with clinical idiopathic NS patient datasets confirmed glomerular ECM dysregulation as a potential mechanism of injury. Cellular deconvolution in silico revealed GC- and Pio-induced amelioration of altered genes primarily within podocytes and mesangial cells. While validation studies are indicated, these analyses identified molecular pathways involved in the early stages of NS (prior to scarring), suggesting that targeting glomerular ECM dysregulation may enable a future non-immunosuppressive approach for proteinuria reduction in idiopathic NS.
Keywords: Gene network; Nephrology; Systems biology; Transcriptomics.
© 2023 The Authors.
Conflict of interest statement
Financial interests: W.E.S. is a co-founder of NephKey Therapeutics, Inc. S.B., S.A., and W.E.S. have filed patent # US 20230257817 A1. Non-financial interests: W.E.S. is on the Board of Directors of NephCure Kidney International and receives no compensation as a member of the Board of Directors.
Figures



Similar articles
-
Nephrotic syndrome-associated hypercoagulopathy is alleviated by both pioglitazone and glucocorticoid which target two different nuclear receptors.Physiol Rep. 2020 Aug;8(15):e14515. doi: 10.14814/phy2.14515. Physiol Rep. 2020. PMID: 32776495 Free PMC article.
-
Pioglitazone enhances proteinuria reduction in complicated pediatric nephrotic syndrome.Pediatr Nephrol. 2023 Apr;38(4):1127-1138. doi: 10.1007/s00467-022-05637-8. Epub 2022 Aug 15. Pediatr Nephrol. 2023. PMID: 35969278
-
Pioglitazone Enhances the Beneficial Effects of Glucocorticoids in Experimental Nephrotic Syndrome.Sci Rep. 2016 May 4;6:24392. doi: 10.1038/srep24392. Sci Rep. 2016. PMID: 27142691 Free PMC article.
-
The impact of steroids on the injured podocytes in nephrotic syndrome.J Steroid Biochem Mol Biol. 2020 Feb;196:105490. doi: 10.1016/j.jsbmb.2019.105490. Epub 2019 Oct 3. J Steroid Biochem Mol Biol. 2020. PMID: 31586640 Review.
-
[Treatment of nephrotic syndrome: immuno- or rather podocyte therapy?].Postepy Hig Med Dosw (Online). 2016 May 5;70:459-70. doi: 10.5604/17322693.1201202. Postepy Hig Med Dosw (Online). 2016. PMID: 27180964 Review. Polish.
Cited by
-
Extracellular Vesicle Mitochondrial DNA Reflects Podocyte Mitochondrial Stress and Is Associated with Relapse in Nephrotic Syndrome.Int J Mol Sci. 2025 Jul 26;26(15):7245. doi: 10.3390/ijms26157245. Int J Mol Sci. 2025. PMID: 40806378 Free PMC article.
References
-
- Trautmann A., Boyer O., Hodson E., Bagga A., Gipson D.S., Samuel S., Wetzels J., Alhasan K., Banerjee S., Bhimma R., et al. IPNA clinical practice recommendations for the diagnosis and management of children with steroid-sensitive nephrotic syndrome. Pediatr. Nephrol. 2023;38:877–919. - PMC - PubMed
-
- McGrogan A., Franssen C.F.M., de Vries C.S. The incidence of primary glomerulonephritis worldwide: a systematic review of the literature. Nephrol. Dial. Transplant. 2011;26:414–430. - PubMed
-
- Mittal P., Agarwal S.K., Singh G., Bhowmik D., Mahajan S., Dinda A., Bagchi S. Spectrum of biopsy-proven renal disease in northern India: A single-centre study. Nephrology. 2020;25:55–62. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

