Identification and Characterization of Jasmonic Acid Methyltransferase Involved in the Formation of Floral Methyl Jasmonate in Hedychium coronarium
- PMID: 38202316
- PMCID: PMC10780636
- DOI: 10.3390/plants13010008
Identification and Characterization of Jasmonic Acid Methyltransferase Involved in the Formation of Floral Methyl Jasmonate in Hedychium coronarium
Abstract
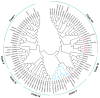
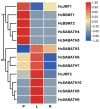
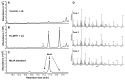
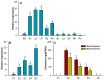
Hedychium coronarium is a popular ornamental flower in tropical and subtropical areas due to its elegant appearance and inviting fragrance. Methyl jasmonate (MeJA) is one of the volatile compounds in the blooming flowers of H. coronarium. However, the molecular mechanism underlying floral MeJA formation is still unclear in H. coronarium. In this study, a total of 12 SABATH family genes were identified in the genome of H. coronarium, and their encoded proteins range from 366 to 387 amino acids. Phylogenetic analysis revealed seven clades in the SABATH family and a JMT ortholog clade, including two HcSABATH members. Combined with expression profiling of HcSABATH members, HcJMT1 was identified as the top candidate gene for floral MeJA biosynthesis. In vitro enzyme assays showed that HcJMT1 can catalyze the production of MeJA from jasmonic acid. Gene expression analysis indicated that HcJMT1 exhibited the highest expression in the labella and lateral petals, the major sites of MeJA emission. During flower development, the two MeJA isomers, major isomers in the products of the HcJMT1 protein, were released after anthesis, in which stage HcJMT1 displayed high expression. Our results indicated that HcJMT1 is involved in the formation of floral MeJA in H. coronarium.
Keywords: Hedychium coronarium; JMT; SABATH methyltransferase; floral volatiles; methyl jasmonate.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Coordinated and High-Level Expression of Biosynthetic Pathway Genes Is Responsible for the Production of a Major Floral Scent Compound Methyl Benzoate in Hedychium coronarium.Front Plant Sci. 2021 Apr 7;12:650582. doi: 10.3389/fpls.2021.650582. eCollection 2021. Front Plant Sci. 2021. PMID: 33897740 Free PMC article.
-
Genome-Wide Analysis Reveals the Potential Role of MYB Transcription Factors in Floral Scent Formation in Hedychium coronarium.Front Plant Sci. 2021 Feb 26;12:623742. doi: 10.3389/fpls.2021.623742. eCollection 2021. Front Plant Sci. 2021. PMID: 33719296 Free PMC article.
-
A Hedychium coronarium short chain alcohol dehydrogenase is a player in allo-ocimene biosynthesis.Plant Mol Biol. 2019 Oct;101(3):297-313. doi: 10.1007/s11103-019-00904-z. Epub 2019 Jul 31. Plant Mol Biol. 2019. PMID: 31368003
-
Phytochemistry and pharmacology of ornamental gingers, Hedychium coronarium and Alpinia purpurata: a review.J Integr Med. 2015 Nov;13(6):368-79. doi: 10.1016/S2095-4964(15)60208-4. J Integr Med. 2015. PMID: 26559362 Review.
-
Methyl jasmonate as a vital substance in plants.Trends Genet. 2003 Jul;19(7):409-13. doi: 10.1016/S0168-9525(03)00138-0. Trends Genet. 2003. PMID: 12850447 Review.
Cited by
-
Plant Volatile Organic Compounds: Revealing the Hidden Interactions.Plants (Basel). 2025 Feb 7;14(4):507. doi: 10.3390/plants14040507. Plants (Basel). 2025. PMID: 40006764 Free PMC article.
-
Comprehensive Analysis of Hormonal Signaling Pathways and Gene Expression in Flesh Segment Development of Chinese Bayberry (Myrica rubra).Plants (Basel). 2025 Feb 13;14(4):571. doi: 10.3390/plants14040571. Plants (Basel). 2025. PMID: 40006830 Free PMC article.
References
-
- Raguso R.A. Wake up and smell the roses: The ecology and evolution of floral scent. Annu. Rev. Ecol. Evol. Syst. 2008;39:549–569. doi: 10.1146/annurev.ecolsys.38.091206.095601. - DOI
Grants and funding
LinkOut - more resources
Full Text Sources

