In Vivo Phosphorylation of the Cytosolic Glucose-6-Phosphate Dehydrogenase Isozyme G6PD6 in Phosphate-Resupplied Arabidopsis thaliana Suspension Cells and Seedlings
- PMID: 38202338
- PMCID: PMC10780934
- DOI: 10.3390/plants13010031
In Vivo Phosphorylation of the Cytosolic Glucose-6-Phosphate Dehydrogenase Isozyme G6PD6 in Phosphate-Resupplied Arabidopsis thaliana Suspension Cells and Seedlings
Abstract
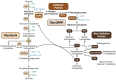
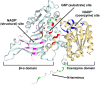
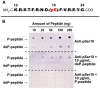
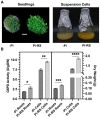
Glucose-6-phosphate dehydrogenase (G6PD) catalyzes the first committed step of the oxidative pentose phosphate pathway (OPPP). Our recent phosphoproteomics study revealed that the cytosolic G6PD6 isozyme became hyperphosphorylated at Ser12, Thr13 and Ser18, 48 h following phosphate (Pi) resupply to Pi-starved (-Pi) Arabidopsis thaliana cell cultures. The aim of the present study was to assess whether G6PD6 phosphorylation also occurs in shoots or roots following Pi resupply to -Pi Arabidopsis seedlings, and to investigate its relationship with G6PD activity. Interrogation of phosphoproteomic databases indicated that N-terminal, multi-site phosphorylation of G6PD6 and its orthologs is quite prevalent. However, the functions of these phosphorylation events remain unknown. Immunoblotting with an anti-(pSer18 phosphosite-specific G6PD6) antibody confirmed that G6PD6 from Pi-resupplied, but not -Pi, Arabidopsis cell cultures or seedlings (i.e., roots) was phosphorylated at Ser18; this correlated with a significant increase in extractable G6PD activity, and biomass accumulation. Peptide kinase assays of Pi-resupplied cell culture extracts indicated that G6PD6 phosphorylation at Ser18 is catalyzed by a Ca2+-dependent protein kinase (CDPK), which correlates with the 'CDPK-like' targeting motif that flanks Ser18. Our results support the hypothesis that N-terminal phosphorylation activates G6PD6 to enhance OPPP flux and thus the production of reducing power (i.e., NADPH) and C-skeletons needed to establish the rapid resumption of growth that ensures Pi-resupply to -Pi Arabidopsis.
Keywords: CDPK; glucose-6-phosphate dehydrogenase; oxidative pentose–phosphate pathway; phosphate nutrition; post-translational modification; protein phosphorylation.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




Similar articles
-
Cytosolic Glucose-6-Phosphate Dehydrogenase Is Involved in Seed Germination and Root Growth Under Salinity in Arabidopsis.Front Plant Sci. 2019 Feb 22;10:182. doi: 10.3389/fpls.2019.00182. eCollection 2019. Front Plant Sci. 2019. PMID: 30873191 Free PMC article.
-
Alternative splicing of Arabidopsis G6PD5 recruits NADPH-producing OPPP reactions to the endoplasmic reticulum.Front Plant Sci. 2022 Sep 2;13:909624. doi: 10.3389/fpls.2022.909624. eCollection 2022. Front Plant Sci. 2022. PMID: 36119606 Free PMC article.
-
Evidence for peroxisomal redundancy among the glucose-6-phosphate dehydrogenase isoforms of Arabidopsis thaliana.Plant Cell Physiol. 2025 May 30;66(5):722-737. doi: 10.1093/pcp/pcaf012. Plant Cell Physiol. 2025. PMID: 39829315 Free PMC article.
-
Loss of cytosolic glucose-6-phosphate dehydrogenase increases the susceptibility of Arabidopsis thaliana to root-knot nematode infection.Ann Bot. 2019 Jan 1;123(1):37-46. doi: 10.1093/aob/mcy124. Ann Bot. 2019. PMID: 29992234 Free PMC article.
-
The regulation of the oxidative phase of the pentose phosphate pathway: new answers to old problems.IUBMB Life. 2014 Nov;66(11):775-9. doi: 10.1002/iub.1329. Epub 2014 Nov 18. IUBMB Life. 2014. PMID: 25408203 Review.
References
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

