Dynamin-Related Protein 1 Binding Partners MiD49 and MiD51 Increased Mitochondrial Fission In Vitro and Atherosclerosis in High-Fat-Diet-Fed ApoE-/- Mice
- PMID: 38203413
- PMCID: PMC10778634
- DOI: 10.3390/ijms25010244
Dynamin-Related Protein 1 Binding Partners MiD49 and MiD51 Increased Mitochondrial Fission In Vitro and Atherosclerosis in High-Fat-Diet-Fed ApoE-/- Mice
Abstract
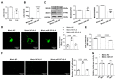
Novel components of the mitochondrial fission machinery, mitochondrial dynamics proteins of 49 kDa (MiD49) and 51 kDa (MiD51), have been recently described, and their potential therapeutic targets for treating cardiovascular disease have been shown, including acute myocardial infarction (AMI), anthracycline cardiomyopathy and pulmonary arterial hypertension (PAH). Here, we examined the role of MiD49 and MiD51 in atherosclerosis. MiD49/51 expression was increased in the aortic valve endothelial cells (ECs) of high-fat diet-induced atherosclerosis in ApoE-/-mice and IL-8-induced human umbilical vein endothelial cells (HUVECs), which accelerated dynamin-related protein 1 (Drp1)-mediated mitochondrial fission. Silencing MiD49/51 reduced atherosclerotic plaque size, increased collagen content, and decreased the IL-8-induced adhesion and proliferation of HUVECs. MiD51 upregulation resulted from decreased microRNA (miR)-107 expression and increased hypoxia-inducible factor-1a (HIF-1a) expression. Treatment with miR-107 mimics decreased atherosclerotic plaque size by reducing HIF-1α and MiD51 production. Both MiD49 and MiD51 were involved in atherosclerotic plaque formation through Drp1-mediated mitochondrial fission, and the involvement of MiD51 in this process was the result of decreased miR-107 expression and increased HIF-1α expression. The miR-107-HIF-1α-MiD51 pathway might provide new therapeutic targets for atherosclerosis.
Keywords: MiD49; MiD51; atherosclerosis; endothelial cells; mitochondrial fission.
Conflict of interest statement
The authors declare no conflict of interest.
Figures








Similar articles
-
Epigenetic Dysregulation of the Dynamin-Related Protein 1 Binding Partners MiD49 and MiD51 Increases Mitotic Mitochondrial Fission and Promotes Pulmonary Arterial Hypertension: Mechanistic and Therapeutic Implications.Circulation. 2018 Jul 17;138(3):287-304. doi: 10.1161/CIRCULATIONAHA.117.031258. Epub 2018 Feb 5. Circulation. 2018. PMID: 29431643 Free PMC article.
-
Adaptor proteins MiD49 and MiD51 can act independently of Mff and Fis1 in Drp1 recruitment and are specific for mitochondrial fission.J Biol Chem. 2013 Sep 20;288(38):27584-27593. doi: 10.1074/jbc.M113.479873. Epub 2013 Aug 6. J Biol Chem. 2013. PMID: 23921378 Free PMC article.
-
The role of Drp1 adaptor proteins MiD49 and MiD51 in mitochondrial fission: implications for human disease.Clin Sci (Lond). 2016 Nov 1;130(21):1861-74. doi: 10.1042/CS20160030. Clin Sci (Lond). 2016. PMID: 27660309 Review.
-
Crystal structure and functional analysis of MiD49, a receptor for the mitochondrial fission protein Drp1.Protein Sci. 2015 Mar;24(3):386-94. doi: 10.1002/pro.2629. Epub 2015 Jan 14. Protein Sci. 2015. PMID: 25581164 Free PMC article.
-
The Role of Mitochondrial Dynamics and Mitotic Fission in Regulating the Cell Cycle in Cancer and Pulmonary Arterial Hypertension: Implications for Dynamin-Related Protein 1 and Mitofusin2 in Hyperproliferative Diseases.Cells. 2023 Jul 20;12(14):1897. doi: 10.3390/cells12141897. Cells. 2023. PMID: 37508561 Free PMC article. Review.
Cited by
-
Cordycepin Ameliorates Renal Interstitial Fibrosis by Inhibiting Drp1-Mediated Mitochondrial Fission.Drug Des Devel Ther. 2025 Feb 24;19:1271-1287. doi: 10.2147/DDDT.S498525. eCollection 2025. Drug Des Devel Ther. 2025. PMID: 40026334 Free PMC article.
-
The perspective of modern transplant science - transplant arteriosclerosis: inspiration derived from mitochondria associated endoplasmic reticulum membrane dysfunction in arterial diseases.Int J Surg. 2025 May 1;111(5):3430-3440. doi: 10.1097/JS9.0000000000002362. Int J Surg. 2025. PMID: 40146783 Free PMC article. Review.
-
Decoding interaction between mitochondria and endoplasmic reticulum in ischemic myocardial injury: targeting natural medicines.Front Pharmacol. 2025 Feb 28;16:1536773. doi: 10.3389/fphar.2025.1536773. eCollection 2025. Front Pharmacol. 2025. PMID: 40093324 Free PMC article. Review.
-
Overexpression of SOX4 in MSCs inhibits cellular senescence and enhances therapeutic efficacy in systemic lupus erythematosus.Stem Cell Res Ther. 2025 Jul 16;16(1):380. doi: 10.1186/s13287-025-04525-w. Stem Cell Res Ther. 2025. PMID: 40671148 Free PMC article.
-
Mitochondrial quality control in diabetes mellitus and complications: molecular mechanisms and therapeutic strategies.Cell Death Dis. 2025 Aug 27;16(1):652. doi: 10.1038/s41419-025-07936-y. Cell Death Dis. 2025. PMID: 40866350 Free PMC article. Review.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

