Genetic diversity and evolutionary patterns of SARS-CoV-2 among the Bhutanese population during the pandemic
- PMID: 38204428
- PMCID: PMC10788421
- DOI: 10.24171/j.phrp.2023.0209
Genetic diversity and evolutionary patterns of SARS-CoV-2 among the Bhutanese population during the pandemic
Abstract
Background: The coronavirus disease 2019 (COVID-19) pandemic, caused by a dynamic virus, has had a profound global impact. Despite declining global COVID-19 cases and mortality rates, the emergence of new severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) variants remains a major concern. This study provides a comprehensive analysis of the genomic sequences of SARS-CoV-2 within the Bhutanese population during the pandemic. The primary aim was to elucidate the molecular epidemiology and evolutionary patterns of SARS-CoV-2 in Bhutan, with a particular focus on genetic variations and lineage dynamics.
Methods: Whole-genome sequences of SARS-CoV-2 collected from Bhutan between May 2020 and February 2023 (n=135) were retrieved from the Global Initiative on Sharing All Influenza Database.
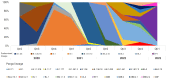
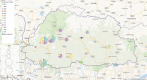
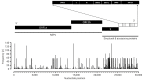
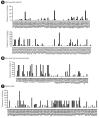
Results: The SARS-CoV-2 variants in Bhutan were predominantly classified within the Nextstrain clade 20A (31.1%), followed by clade 21L (20%) and clade 22D (15.6%). We identified 26 Pangolin lineages with variations in their spatial and temporal distribution. Bayesian time-scaled phylogenetic analysis estimated the time to the most recent common ancestor as February 15, 2020, with a substitution rate of 0.97×10-3 substitutions per site per year. Notably, the spike glycoprotein displayed the highest mutation frequency among major viral proteins, with 116 distinct mutations, including D614G. The Bhutanese isolates also featured mutations such as E484K, K417N, and S477N in the spike protein, which have implications for altered viral properties.
Conclusion: This is the first study to describe the genetic diversity of SARS-CoV-2 circulating in Bhutan during the pandemic, and this data can inform public health policies and strategies for preventing future outbreaks in Bhutan.
Keywords: Coronavirus spike glycoprotein; Molecular epidemiology; Mutation; SARS-CoV-2.
Conflict of interest statement
The authors have no conflicts of interest to declare.
Figures



Similar articles
-
Genetic diversity and genomic epidemiology of SARS-CoV-2 during the first 3 years of the pandemic in Morocco: comprehensive sequence analysis, including the unique lineage B.1.528 in Morocco.Access Microbiol. 2024 Oct 7;6(10):000853.v4. doi: 10.1099/acmi.0.000853.v4. eCollection 2024. Access Microbiol. 2024. PMID: 39376591 Free PMC article.
-
Genome sequence diversity of SARS-CoV-2 in Serbia: insights gained from a 3-year pandemic study.Front Microbiol. 2024 Feb 27;15:1332276. doi: 10.3389/fmicb.2024.1332276. eCollection 2024. Front Microbiol. 2024. PMID: 38476954 Free PMC article.
-
A Founder Effect Led Early SARS-CoV-2 Transmission in Spain.J Virol. 2021 Jan 13;95(3):e01583-20. doi: 10.1128/JVI.01583-20. Print 2021 Jan 13. J Virol. 2021. PMID: 33127745 Free PMC article.
-
Molecular evolution of SARS-CoV-2 from December 2019 to August 2022.J Med Virol. 2023 Jan;95(1):e28366. doi: 10.1002/jmv.28366. J Med Virol. 2023. PMID: 36458547 Free PMC article. Review.
-
The Development of SARS-CoV-2 Variants: The Gene Makes the Disease.J Dev Biol. 2021 Dec 15;9(4):58. doi: 10.3390/jdb9040058. J Dev Biol. 2021. PMID: 34940505 Free PMC article. Review.
References
-
- World Health Organization (WHO). Statement on the fifteenth meeting of the IHR (2005) Emergency Committee on the COVID-19 pandemic [Internet]. WHO; 2023 [cited 2023 May 18]. Available from: https://www.who.int/news/item/05-05-2023-statement-on-the-fifteenth-meet....
-
- World Health Organization (WHO). WHO coronavirus (COVID-19) dashboard [Internet]. WHO; 2023 [cited 2023 May 18]. Available from: https://covid19.who.int/
LinkOut - more resources
Full Text Sources
Miscellaneous

