Predicting the final metabolic profile based on the succession-related microbiota during spontaneous fermentation of the starter for Chinese liquor making
- PMID: 38206013
- PMCID: PMC10878095
- DOI: 10.1128/msystems.00586-23
Predicting the final metabolic profile based on the succession-related microbiota during spontaneous fermentation of the starter for Chinese liquor making
Abstract
Microbial inoculation is an effective way to improve the quality of fermented foods via affecting the microbiota structure. However, it is unclear how the inoculation regulates the microbiota structure, and it is still difficult to directionally control the microbiota function via the inoculation. In this work, using the spontaneous fermentation of the starter (Daqu) for Chinese liquor fermentation as a case, we inoculated different microbiota groups at different time points in Daqu fermentation, and analyzed the effect of the inoculation on the final metabolic profile of Daqu. The inoculated microbiota and inoculated time points both significantly affected the final metabolites via regulating the microbial succession (P < 0.001), and multiple inoculations can promote deterministic assembly. Twenty-seven genera were identified to be related to microbial succession, and drove the variation of 121 metabolites. We then constructed an elastic network model to predict the profile of these 121 metabolites based on the abundances of 27 succession-related genera in Daqu fermentation. Procrustes analysis showed that the model could accurately predict the metabolic abundances (average Spearman correlation coefficients >0.3). This work revealed the effect of inoculation on the microbiota succession and the metabolic profile. The established predicted model of metabolic profile would be beneficial for directionally improving the food quality.IMPORTANCEThis work revealed the importance of microbial succession to microbiota structure and metabolites. Multi-inoculations would promote deterministic assembly. It would facilitate the regulation of microbiota structure and metabolic profile. In addition, we established a model to predict final metabolites based on microbial genera related to microbial succession. This model was beneficial for optimizing the inoculation of the microbiota. This work would be helpful for controlling the spontaneous food fermentation and directionally improving the food quality.
Keywords: liquor fermentation; metabolites; microbial succession; microbiota; modeling.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


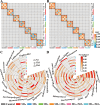


References
-
- Blandino A, Al-Aseeri ME, Pandiella SS, Cantero D, Webb C. 2003. Cereal-based fermented foods and beverages. Food Res Internat 36:527–543. doi:10.1016/S0963-9969(03)00009-7 - DOI
-
- Liu S, Yang L, Zhou Y, He S, Li J, Sun H, Yao S, Xu S. 2019. Effect of mixed moulds starters on volatile flavor compounds in rice wine. LWT 112:108215. doi:10.1016/j.lwt.2019.05.113 - DOI
-
- Isaac Newton Institute Fellows, Widder S, Allen RJ, Pfeiffer T, Curtis TP, Wiuf C, Sloan WT, Cordero OX, Brown SP, Momeni B, et al. . 2016. Challenges in microbial ecology: building predictive understanding of community function and dynamics. ISME J 10:2557–2568. doi:10.1038/ismej.2016.45 - DOI - PMC - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
